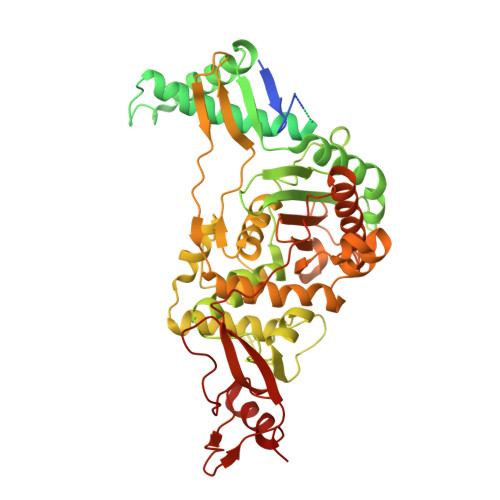
Structural and Mechanistic Basis of Penicillin-Binding Protein Inhibition by Lactivicins
Macheboeuf, P., Fisher, D.S., Brown, T.J., Zervosen, A., Luxen, A., Joris, B., Dessen, A., Schofield, C.J.(2007) Nat Chem Biol 3: 565
- PubMed: 17676039
- DOI: https://doi.org/10.1038/nchembio.2007.21
- Primary Citation of Related Structures:
2JCH, 2JE5 - PubMed Abstract:
Beta-lactam antibiotics, including penicillins and cephalosporins, inhibit penicillin-binding proteins (PBPs), which are essential for bacterial cell wall biogenesis. Pathogenic bacteria have evolved efficient antibiotic resistance mechanisms that, in Gram-positive bacteria, include mutations to PBPs that enable them to avoid beta-lactam inhibition. Lactivicin (LTV; 1) contains separate cycloserine and gamma-lactone rings and is the only known natural PBP inhibitor that does not contain a beta-lactam. Here we show that LTV and a more potent analog, phenoxyacetyl-LTV (PLTV; 2), are active against clinically isolated, penicillin-resistant Streptococcus pneumoniae strains. Crystallographic analyses of S. pneumoniae PBP1b reveal that LTV and PLTV inhibition involves opening of both monocyclic cycloserine and gamma-lactone rings. In PBP1b complexes, the ring-derived atoms from LTV and PLTV show a notable structural convergence with those derived from a complexed cephalosporin (cefotaxime; 3). The structures imply that derivatives of LTV will be useful in the search for new antibiotics with activity against beta-lactam-resistant bacteria.
Organizational Affiliation:
Institut de Biologie Structurale Jean-Pierre Ebel Commissariat à l'énergie atomique - Centre National de La Recherche Scientifique - Université Joseph Fourier, 41 rue Jules Horowitz, F-38027 Grenoble, France.