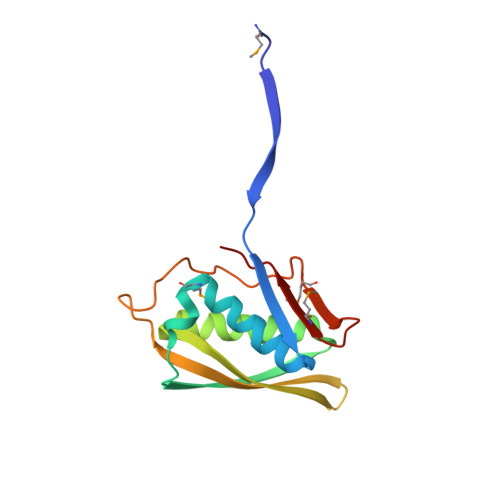
A tRNA splicing operon: Archease endows RtcB with dual GTP/ATP cofactor specificity and accelerates RNA ligation.
Desai, K.K., Cheng, C.L., Bingman, C.A., Phillips Jr., G.N., Raines, R.T.(2014) Nucleic Acids Res 42: 3931-3942
- PubMed: 24435797
- DOI: https://doi.org/10.1093/nar/gkt1375
- Primary Citation of Related Structures:
4N2P - PubMed Abstract:
Archease is a 16-kDa protein that is conserved in all three domains of life. In diverse bacteria and archaea, the genes encoding Archease and the tRNA ligase RtcB are localized into an operon. Here we provide a rationale for this operon organization by showing that Archease and RtcB from Pyrococcus horikoshii function in tandem, with Archease altering the catalytic properties of the RNA ligase. RtcB catalyzes the GTP and Mn(II)-dependent joining of either 2',3'-cyclic phosphate or 3'-phosphate termini to 5'-hydroxyl termini. We find that catalytic concentrations of Archease are sufficient to activate RtcB, and that Archease accelerates both the RNA 3'-P guanylylation and ligation steps. In addition, we show that Archease can alter the NTP specificity of RtcB such that ATP, dGTP or ITP is used efficiently. Moreover, RtcB variants that have inactivating substitutions in the guanine-binding pocket can be rescued by the addition of Archease. We also present a 1.4 Å-resolution crystal structure of P. horikoshii Archease that reveals a metal-binding site consisting of conserved carboxylates located at the protein tip. Substitution of the Archease metal-binding residues drastically reduced Archease-dependent activation of RtcB. Thus, evolution has sought to co-express archease and rtcB by creating a tRNA splicing operon.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin-Madison, Madison, WI 53706, USA, Department of Biochemistry and Cell Biology and Department of Chemistry, Rice University, Houston, TX 77005, USA and Department of Chemistry, University of Wisconsin-Madison, Madison, WI 53706, USA.