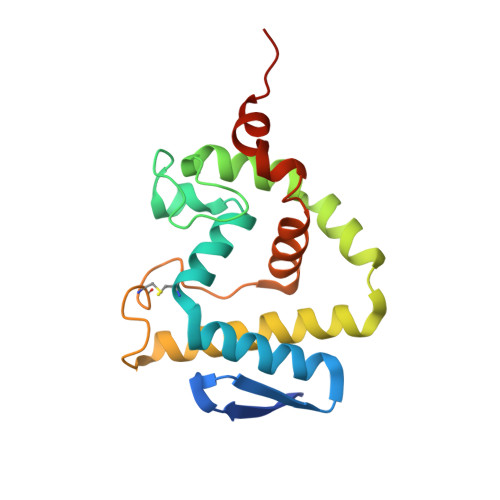
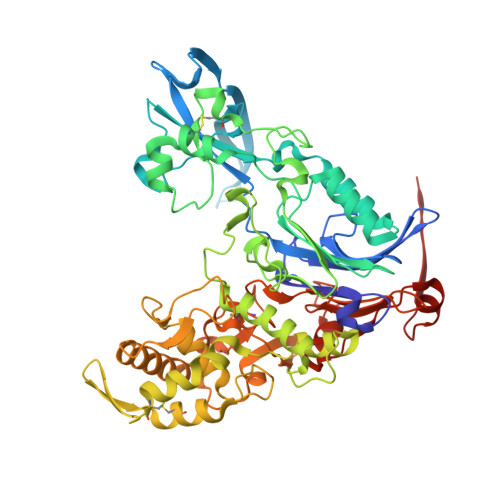
Bifunctional quorum-quenching and antibiotic-acylase MacQ forms a 170-kDa capsule-shaped molecule containing spacer polypeptides
Yasutake, Y., Kusada, H., Ebuchi, T., Hanada, S., Kamagata, Y., Tamura, T., Kimura, N.(2017) Sci Rep 7: 8946-8946
- PubMed: 28827579
- DOI: https://doi.org/10.1038/s41598-017-09399-4
- Primary Citation of Related Structures:
4YF9, 4YFA, 4YFB, 5C9I - PubMed Abstract:
Understanding the molecular mechanisms of bacterial antibiotic resistance will help prepare against further emergence of multi-drug resistant strains. MacQ is an enzyme responsible for the multi-drug resistance of Acidovorax sp. strain MR-S7. MacQ has acylase activity against both N-acylhomoserine lactones (AHLs), a class of signalling compounds involved in quorum sensing, and β-lactam antibiotics. Thus, MacQ is crucial as a quencher of quorum sensing as well as in conferring antibiotic resistance in Acidovorax. Here, we report the X-ray structures of MacQ in ligand-free and reaction product complexes. MacQ forms a 170-kDa capsule-shaped molecule via face-to-face interaction with two heterodimers consisting of an α-chain and a β-chain, generated by the self-cleaving activity of a precursor polypeptide. The electron density of the spacer polypeptide in the hollow of the molecule revealed the close orientation of the peptide-bond atoms of Val20SP-Gly21SP to the active-site, implying a role of the residues in substrate binding. In mutational analyses, uncleaved MacQ retained degradation activity against both AHLs and penicillin G. These results provide novel insights into the mechanism of self-cleaving maturation and enzymatic function of N-terminal nucleophile hydrolases.
Organizational Affiliation:
Bioproduction Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), 2-17-2-1 Tsukisamu-Higashi, Toyohira, Sapporo, 062-8517, Japan. [email protected].