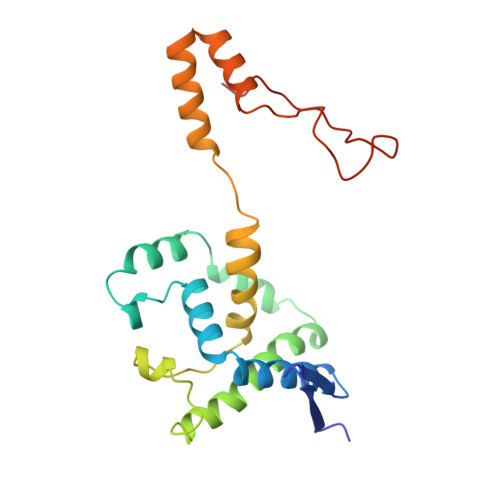
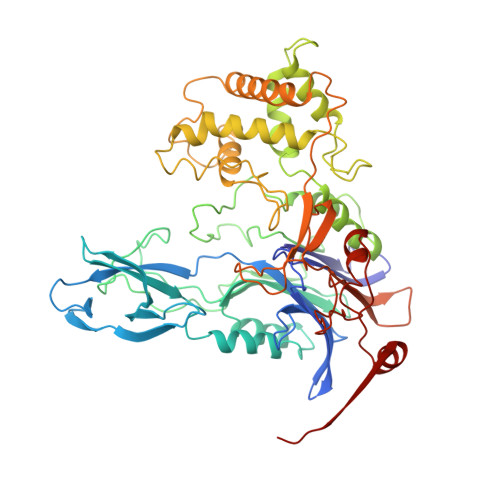
Ligand-induced conformational change in penicillin acylase.
Done, S.H., Brannigan, J.A., Moody, P.C.E., Hubbard, R.E.(1998) J Mol Biol 284: 463-475
- PubMed: 9813130
- DOI: https://doi.org/10.1006/jmbi.1998.2180
- Primary Citation of Related Structures:
1AI4, 1AI5, 1AI6, 1AI7, 1AJN, 1AJP, 1AJQ - PubMed Abstract:
The enzyme penicillin acylase (penicillin amidohydrolase EC 3.5.1. 11) catalyses the cleavage of the amide bond in the benzylpenicillin (penicillin G) side-chain to produce phenylacetic acid and 6-aminopenicillanic acid (6-APA). The enzyme is of great pharmaceutical importance, as the product 6-APA is the starting point for the synthesis of many semi-synthetic penicillin antibiotics. Studies have shown that the enzyme is specific for hydrolysis of phenylacetamide derivatives, but is more tolerant of features in the rest of the substrate. It is this property that has led to many other applications for the enzyme, and greater knowledge of the enzyme's structure and specificity could facilitate engineering of the enzyme, enhancing its potential for chemical and industrial applications. An extensive study of the binding of a series of phenylacetic acid derivatives has been carried out. A measure of the relative degree of inhibition of the enzyme by each of the compounds has been obtained using a competitive inhibition assay, and the structures of a number of these complexes have been determined by X-ray crystallography. The structures reveal a clear rationale for the observed kinetic results, but show also that some of the ligands cause a conformational change within the binding pocket. This change can generally be understood in terms of the size and orientation of the ligand within the active site.The results reveal that ligand binding in penicillin acylase is facilitated by certain amino acid residues that can adopt two distinct, energetically favourable positions in order to accommodate a variety of compounds within the active site. The structures of these complexes provide evidence for conformational changes in the substrate-binding region that may act as a switch in the mechanism of autocatalytic processing of this enzyme.
Organizational Affiliation:
Department of Chemistry, University of York, Heslington, YO1 5DD, UK. [email protected]