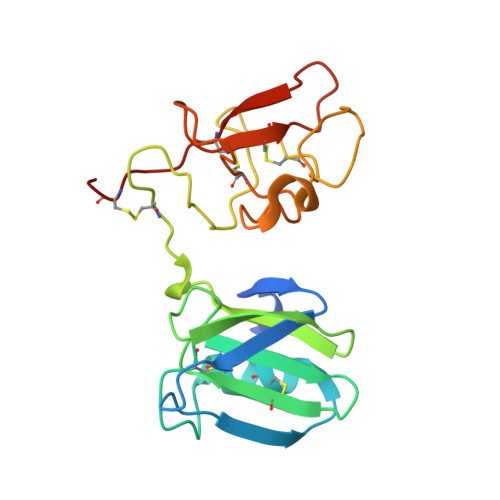
Crystal Structures of Nk1-Heparin Complexes Reveal the Basis for Nk1 Activity and Enable Engineering of Potent Agonists of the met Receptor
Lietha, D., Chirgadze, D.Y., Mulloy, B., Blundell, T.L., Gherardi, E.(2001) EMBO J 20: 5543
- PubMed: 11597998
- DOI: https://doi.org/10.1093/emboj/20.20.5543
- Primary Citation of Related Structures:
1GMN, 1GMO - PubMed Abstract:
NK1 is a splice variant of the polypeptide growth factor HGF/SF, which consists of the N-terminal (N) and first kringle (K) domain and requires heparan sulfate or soluble heparin for activity. We describe two X-ray crystal structures of NK1-heparin complexes that define a heparin-binding site in the N domain, in which a major role is played by R73, with further contributions from main chain atoms of T61, K63 and G79 and the side chains of K60, T61, R76, K62 and K58. Mutagenesis experiments demonstrate that heparin binding to this site is essential for dimerization in solution and biological activity of NK1. Heparin also comes into contact with a patch of positively charged residues (K132, R134, K170 and R181) in the K domain. Mutation of these residues yields NK1 variants with increased biological activity. Thus, we uncover a complex role for heparan sulfate in which binding to the primary site in the N domain is essential for biological activity whereas binding to the K domain reduces activity. We exploit the interaction between heparin and the K domain site in order to engineer NK1 as a potent receptor agonist and suggest that dual (positive and negative) control may be a general mechanism of heparan sulfate-dependent regulation of growth factor activity.
Organizational Affiliation:
Growth Factors Group, MRC Centre, Hills Road, Cambridge CB2 2QH, UK.