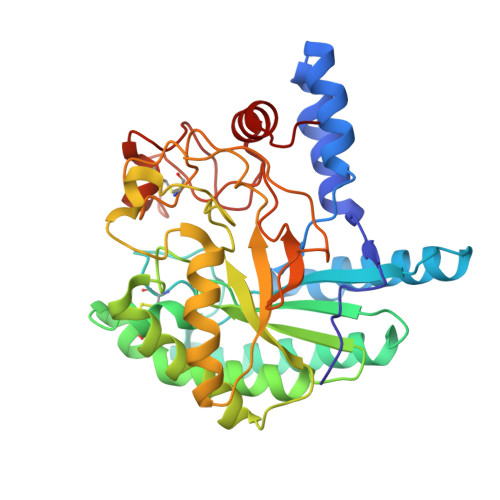
Structure of the Humicola Insolens Cellobiohydrolase Cel6A D416A Mutant in Complex with a Non-Hydrolysable Substrate Analogue, Methyl Cellobiosyl-4-Thio-Beta-Cellobioside, at 1.9 A
Varrot, A., Frandsen, T., Driguez, H., Davies, G.J.(2002) Acta Crystallogr D Biol Crystallogr 58: 2201
- PubMed: 12454501
- DOI: https://doi.org/10.1107/s0907444902017006
- Primary Citation of Related Structures:
1GZ1 - PubMed Abstract:
The enzymatic degradation of cellulose continues to be one of the most important enzyme-catalysed reactions. Glycoside hydrolases from family GH-6 hydrolyse cellulose with inversion of the configuration of the anomeric carbon. Whilst the catalytic proton donor has been clearly identified (Asp226 in Humicola insolens Cel6A), the identification and even the existence of a potential Brønsted base remains unclear. Equally controversial is the role of surface-loop flexibility. Here, the structure of the D416A mutant of the H. insolens cellobiohydrolase Cel6A in complex with a non-hydrolysable thiooligosaccharide methyl cellobiosyl-4-thio-beta-cellobioside at 1.9 A resolution is presented. Substrate distortion in the -1 subsite, to a (2)S(0) skew-boat conformation, is observed, similar to that seen in the analogous Trichoderma reesei Cel6A structure [Zou et al. (1999), Structure, 7, 1035-1045], but the active-centre N-terminal loop of the H. insolens enzyme is found in a more open conformation than described for previous structures.
Organizational Affiliation:
Structural Biology Laboratory, Department of Chemistry, The University of York, Heslington, York Y010 5YW, England.