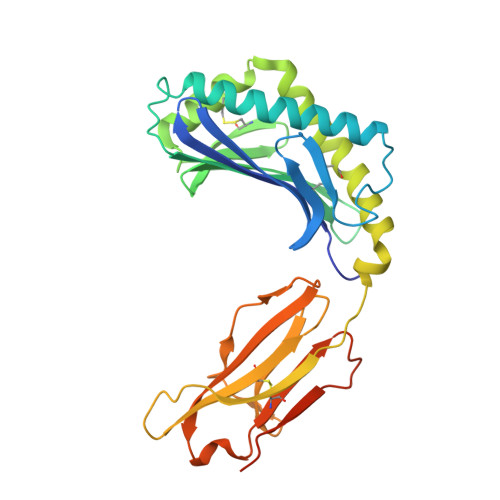
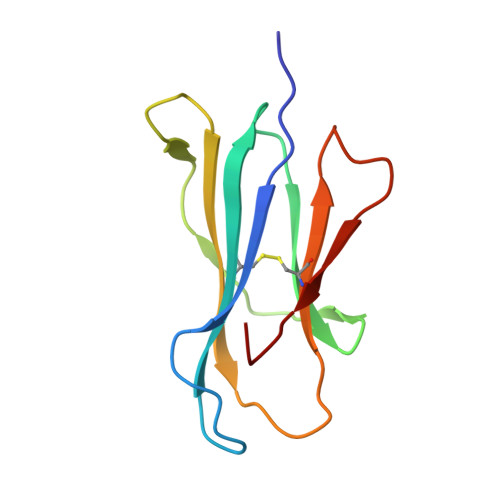
Structure of human CD1b with bound ligands at 2.3 A, a maze for alkyl chains.
Gadola, S.D., Zaccai, N.R., Harlos, K., Shepherd, D., Castro-Palomino, J.C., Ritter, G., Schmidt, R.R., Jones, E.Y., Cerundolo, V.(2002) Nat Immunol 3: 721-726
- PubMed: 12118248
- DOI: https://doi.org/10.1038/ni821
- Primary Citation of Related Structures:
1GZP, 1GZQ - PubMed Abstract:
The human genome encodes five nonpolymorphic major histocompatibility complex class I-like glycoproteins, CD1a to CD1e, that present lipid antigens for specific recognition by T lymphocytes. Using single alkyl chain detergents, we developed a protocol to generate recombinant human CD1b-lipid complexes. We present here the crystal structures of CD1b in complex with either phosphatidylinositol or ganglioside GM2 at 2.3 A and 2.8 A resolutions, respectively. The antigen-binding groove houses four interlinked hydrophobic channels that are occupied by the alkyl chains of the glycolipid plus two detergent molecules. A distinct exit beneath the alpha 2 helix further contributes to the plasticity of the binding groove. These structures reveal the mechanism by which two alkyl chain lipids bind to CD1b, and how CD1b can adapt to ligands of different alkyl chain length. They also suggest how very long alkyl chains, such as those of mycolic acid, could be fully contained within the binding groove. These results extend the spectrum of potential CD1b ligands by revealing that, in addition to two alkyl chain lipids, mono-alkyl and triple-alkyl chain lipids can be accommodated in the binding groove.
Organizational Affiliation:
Cancer Research UK Tumour Immunology Group, The Weatherall Institute of Molecular Medicine, Nuffield Department of Medicine, University of Oxford, Oxford OX3 9DS, UK. stephan.gadola@inselch