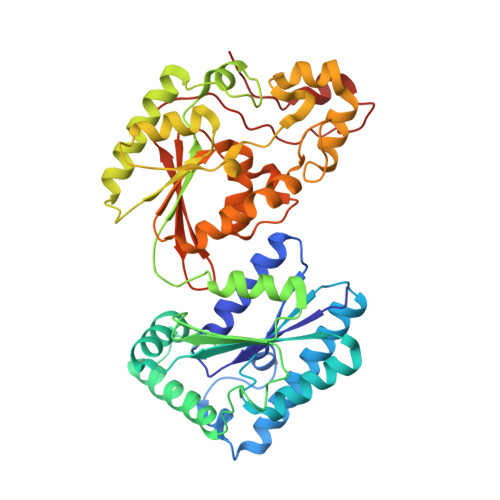
Tissue-specific structure/function differentiation of the liver isoform of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase.
Lee, Y.H., Li, Y., Uyeda, K., Hasemann, C.A.(2003) J Biol Chem 278: 523-530
- PubMed: 12379646
- DOI: https://doi.org/10.1074/jbc.M209105200
- Primary Citation of Related Structures:
1K6M - PubMed Abstract:
The crystal structures of the human liver 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase in three different liganding states were determined and compared with those of the rat testis isozyme. A set of amino acid sequence heterogeneity from the two distinct genes encoding the two different tissue isozymes leads to both global and local conformational differences that may cause the differences in catalytic properties of the two isozymes. The sequence differences in a beta-hairpin loop in the kinase domain causes a translational shift of several hydrophobic interactions in the dimeric contact region, and its propagation to the domains interface results in a 5 degrees twist of the entire bisphosphatase domain relative to the kinase domain. The bisphosphatase domain twist allows the dimeric interactions between the bisphosphatase domains, which are negligible in the testis enzyme, and as a result, the conformational stability of the domain is increased. Sequence polymorphisms also confer small but significant structural dissimilarities in the substrate-binding loops, allowing the differentiated catalytic properties between the two different tissue-type isozymes. Whereas the polymorphic sequence at the bisphosphatase-active pocket suggests a more suitable substrate binding, a similar extent of sequence differences at the kinase-active pocket confers a different mechanism of substrates bindings to the kinase-active pocket. It includes the ATP-sensitive unwinding of the switch helix alpha5, which is a characteristic ATP-dependent conformational change in the testis form. The sequence-dependent structural difference disallows the liver kinase to follow the ATP-switch mechanism. Altogether these suggest that the liver isoform has structural features more appropriate for an elevated bisphosphatase activity, compared with that of the testis form. The structural predisposition for bisphosphatase activity in the liver isozyme is consistent with the liver-unique glucose metabolic pathway, gluconeogenesis.
Organizational Affiliation:
Structural Biology Core, Molecular Biology, University of Missouri, Columbia, Missouri 65211, USA. [email protected]