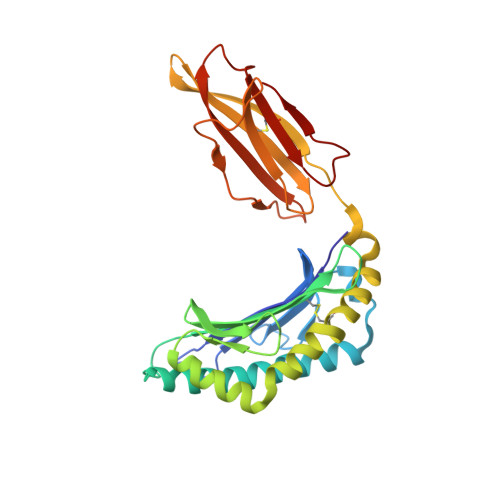
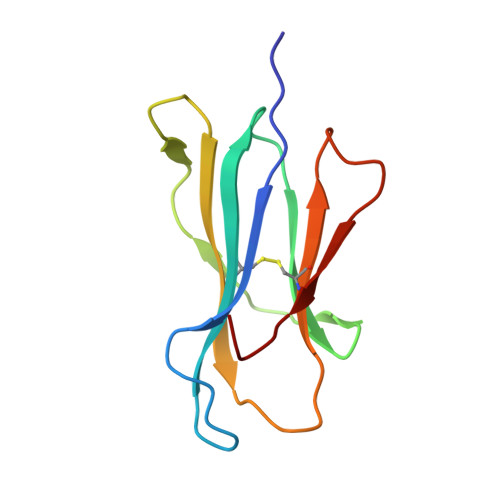
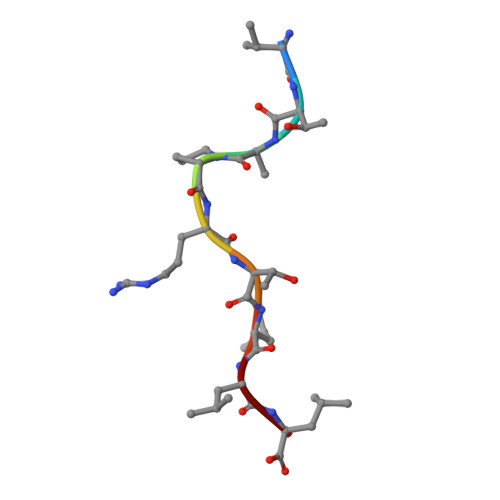
HLA-E allelic variants: Correlating differential expression, peptide affinities, crystal structures and thermal stabilities
Strong, R.K., Holmes, M.A., Li, P., Braun-Jones, L., Lee, N., Geraghty, D.E.(2003) J Biol Chem 278: 5082-5089
- PubMed: 12411439
- DOI: https://doi.org/10.1074/jbc.M208268200
- Primary Citation of Related Structures:
1KPR, 1KTL - PubMed Abstract:
Previous studies of HLA-E allelic polymorphism have indicated that balancing selection may be acting to maintain two major alleles in most populations, indicating that a functional difference may exist between the alleles. The alleles differ at only one amino acid position, where an arginine at position 107 in HLA-E*0101 (E(R)) is replaced by a glycine in HLA-E*0103 (E(G)). To investigate possible functional differences, we have undertaken a study of the physical and biochemical properties of these two proteins. By comparing expression levels, we found that whereas steady-state protein levels were similar, the two alleles did in fact differ with respect to cell surface levels. To help explain this difference, we undertook studies of the relative differences in peptide affinity, complex stability, and three-dimensional structure between the alleles. The crystal structures for HLA-E(G) complexed with two distinct peptides were determined, and both were compared with the HLA-E(R) structure. No significant differences in the structure of HLA-E were induced as a result of binding different peptides or by the allelic substitution at position 107. However, there were clear differences in the relative affinity for peptide of each heavy chain, which correlated with and may be explained by differences between their thermal stabilities. These differences were completely consistent with the relative levels of the HLA-E alleles on the cell surface and may indeed correlate with functional differences. This in turn may help explain the apparent balancing selection acting on this locus.
Organizational Affiliation:
Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA. [email protected]