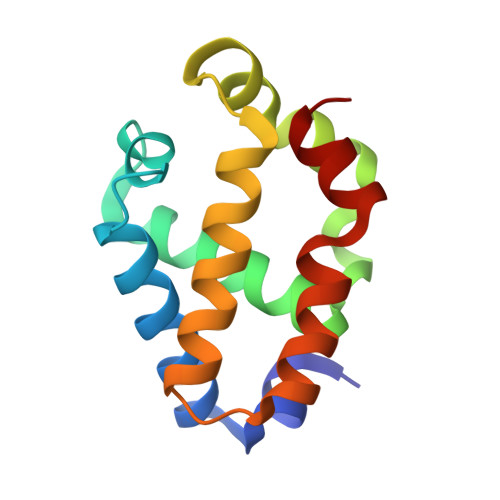
The crystal structure of synechocystis hemoglobin with a covalent heme linkage.
Hoy, J.A., Kundu, S., Trent, J.T., Ramaswamy, S., Hargrove, M.S.(2004) J Biol Chem 279: 16535-16542
- PubMed: 14736872
- DOI: https://doi.org/10.1074/jbc.M313707200
- Primary Citation of Related Structures:
1RTX - PubMed Abstract:
The x-ray crystal structure of Synechocystis hemoglobin has been solved to a resolution of 1.8 A. The conformation of this structure is surprisingly different from that of the previously reported solution structure, probably due in part to a covalent linkage between the heme 2-vinyl and His117 that is present in the crystal structure but not in the structure solved by NMR. Synechocystis hemoglobin is a hexacoordinate hemoglobin in which the heme iron is coordinated by both the proximal and distal histidines. It is also a member of the "truncated hemoglobin" family that is much shorter in primary structure than vertebrate and plant hemoglobins. In contrast to other truncated hemoglobins, the crystal structure of Synechocystis hemoglobin displays no "ligand tunnel" and shows that several important amino acid side chains extrude into the solvent instead of residing inside the heme pocket. The stereochemistry of hexacoordination is compared with other hexacoordinate hemoglobins and cytochromes in an effort to illuminate factors contributing to ligand affinity in hexacoordinate hemoglobins.
Organizational Affiliation:
Department of Biochemistry, Biophysics, and Molecular Biology, Iowa State University, Ames, Iowa 50011, USA.