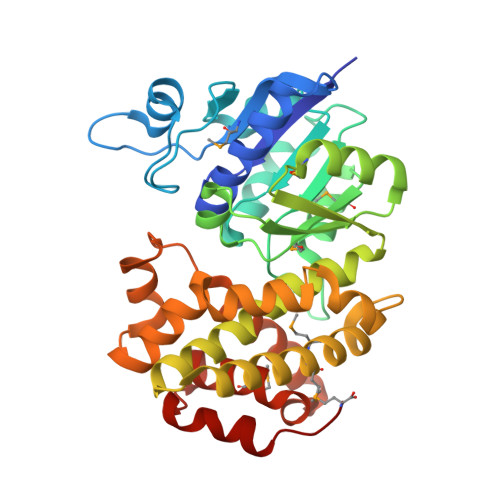
Crystal Structures of Human Glycerol 3-phosphate Dehydrogenase 1 (GPD1)
Ou, X., Ji, C., Han, X., Zhao, X., Li, X., Mao, Y., Wong, L.L., Bartlam, M., Rao, Z.(2006) J Mol Biol 357: 858-869
- PubMed: 16460752
- DOI: https://doi.org/10.1016/j.jmb.2005.12.074
- Primary Citation of Related Structures:
1WPQ, 1X0V, 1X0X - PubMed Abstract:
Homo sapiens L-alpha-glycerol-3-phosphate dehydrogenase 1 (GPD1) catalyzes the reversible biological conversion of dihydroxyacetone (DHAP) to glycerol-3-phosphate. The GPD1 protein was expressed in Escherichia coli, and purified as a fusion protein with glutathione S-transferase. Here we report the apoenzyme structure of GPD1 determined by multiwavelength anomalous diffraction phasing, and other complex structures with small molecules (NAD+ and DHAP) by the molecular replacement method. This enzyme structure is organized into two distinct domains, the N-terminal eight-stranded beta-sheet sandwich domain and the C-terminal helical substrate-binding domain. An electrophilic catalytic mechanism by the epsilon-NH3+ group of Lys204 is proposed on the basis of the structural analyses. In addition, the inhibitory effects of zinc and sulfate on GPDHs are assayed and discussed.
Organizational Affiliation:
National Laboratory of Biomacromolecules, Institute of Biophysics (IBP), Chinese Academy of Sciences, Beijing 100101, China.