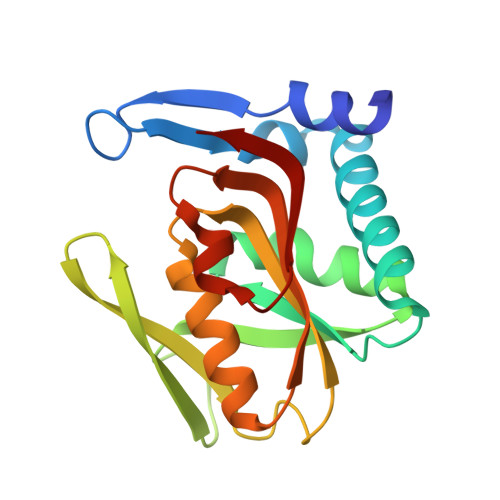
Structural Complexes of Human Adenine Phosphoribosyltransferase Reveal Novel Features of the APRT Catalytic Mechanism
Silva, C.H., Silva, M., Iulek, J., Thiemann, O.H.(2008) J Biomol Struct Dyn 25: 589-598
- PubMed: 18399692
- DOI: https://doi.org/10.1080/07391102.2008.10507205
- Primary Citation of Related Structures:
1ZN7, 1ZN8, 1ZN9 - PubMed Abstract:
Adenine phosphoribosyltransferase (APRT) is an important enzyme component of the purine recycling pathway. Parasitic protozoa of the order Kinetoplastida are unable to synthesize purines de novo and use the salvage pathway for the synthesis of purine bases rendering this biosynthetic pathway an attractive target for antiparasitic drug design. The recombinant human adenine phosphoribosyltransferase (hAPRT) structure was resolved in the presence of AMP in the active site to 1.76 A resolution and with the substrates PRPP and adenine simultaneously bound to the catalytic site to 1.83 A resolution. An additional structure was solved containing one subunit of the dimer in the apo-form to 2.10 A resolution. Comparisons of these three hAPRT structures with other 'type I' PRTases revealed several important features of this class of enzymes. Our data indicate that the flexible loop structure adopts an open conformation before and after binding of both substrates adenine and PRPP. Comparative analyses presented here provide structural evidence to propose the role of Glu104 as the residue that abstracts the proton of adenine N9 atom before its nucleophilic attack on the PRPP anomeric carbon. This work leads to new insights to the understanding of the APRT catalytic mechanism.
Organizational Affiliation:
Departamento de Física e Informática, Grupo de Cristalografia de Proteínas e Biologia Estrutural, Instituto de Física de São Carlos, USP, Caixa Postal 369, 13560-590, São Carlos-SP, Brazil. [email protected]