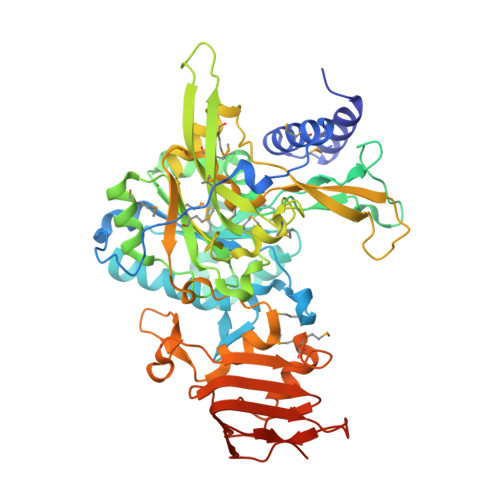
Open and Closed Structures of the UDP-glucose Pyrophosphorylase from Leishmania major.
Steiner, T., Lamerz, A.C., Hess, P., Breithaupt, C., Krapp, S., Bourenkov, G., Huber, R., Gerardy-Schahn, R., Jacob, U.(2007) J Biol Chem 282: 13003-13010
- PubMed: 17303565
- DOI: https://doi.org/10.1074/jbc.M609984200
- Primary Citation of Related Structures:
2OEF, 2OEG - PubMed Abstract:
Uridine diphosphate-glucose pyrophosphorylase (UGPase) represents a ubiquitous enzyme, which catalyzes the formation of UDP-glucose, a key metabolite of the carbohydrate pathways of all organisms. In the protozoan parasite Leishmania major, which causes a broad spectrum of diseases and is transmitted to humans by sand fly vectors, UGPase represents a virulence factor because of its requirement for the synthesis of cell surface glycoconjugates. Here we present the crystal structures of the L. major UGPase in its uncomplexed apo form (open conformation) and in complex with UDP-glucose (closed conformation). The UGPase consists of three distinct domains. The N-terminal domain exhibits species-specific differences in length, which might permit distinct regulation mechanisms. The central catalytic domain resembles a Rossmann-fold and contains key residues that are conserved in many nucleotidyltransferases. The C-terminal domain forms a left-handed parallel beta-helix (LbetaH), which represents a rarely observed structural element. The presented structures together with mutagenesis analyses provide a basis for a detailed analysis of the catalytic mechanism and for the design of species-specific UGPase inhibitors.
Organizational Affiliation:
Max-Planck-Institut für Biochemie, Abteilung für Strukturforschung, Am Klopferspitz 18, 82152 Martinsried, Germany. [email protected]