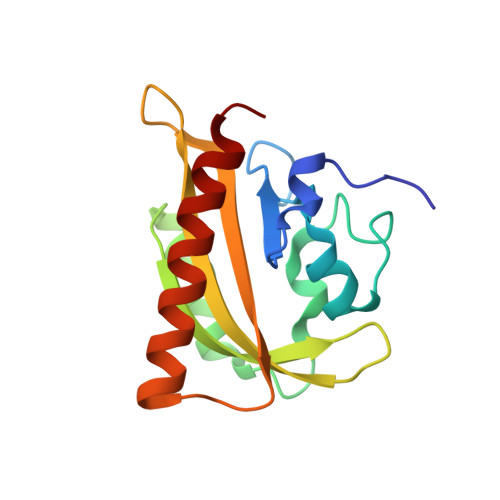
N- and C-Terminal Flanking Regions Modulate Light-Induced Signal Transduction in the Lov2 Domain of the Blue Light Sensor Phototropin 1 from Avena Sativa.
Halavaty, A.S., Moffat, K.(2007) Biochemistry 46: 14001
- PubMed: 18001137
- DOI: https://doi.org/10.1021/bi701543e
- Primary Citation of Related Structures:
2V0U, 2V0W, 2V1A, 2V1B - PubMed Abstract:
Light sensing by photoreceptors controls phototropism, chloroplast movement, stomatal opening, and leaf expansion in plants. Understanding the molecular mechanism by which these processes are regulated requires a quantitative description of photoreceptor dynamics. We focus on a light-driven signal transduction mechanism in the LOV2 domain (LOV, light, oxygen, voltage) of the blue light photoreceptor phototropin 1 from Avena sativa (oat). High-resolution crystal structures of the dark and light states of an oat LOV2 construct including residues Leu404 through Leu546 (LOV2 (404-546)) have been determined at 105 and 293 K. In all four structures, LOV2 (404-546) exhibits the typical Per-ARNT-Sim (PAS) fold, flanked by an additional conserved N-terminal turn-helix-turn motif and a C-terminal flanking region containing an amphipathic Jalpha helix. These regions dock on the LOV2 core domain and bury several hydrophobic residues of the central beta-sheet of the core domain that would otherwise be exposed to solvent. Light structures of LOV2 (404-546) reveal that formation of the covalent bond between Cys450 and the C4a atom of the flavin mononucleotide (FMN) results in local rearrangement of the hydrogen-bonding network in the FMN binding pocket. These rearrangements are associated with disruption of the Asn414-Asp515 hydrogen bond on the surface of the protein and displacement of the N- and C-terminal flanking regions of LOV2 (404-546), both of which constitute a structural signal.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, The University of Chicago, 929 East 57th Street, Chicago, Illinois 60637, USA.