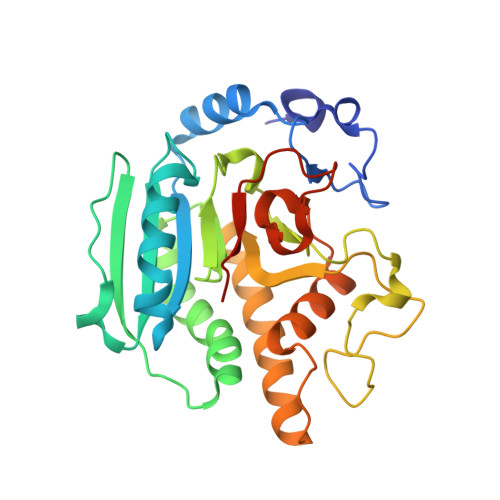
Conformational Changes Induced by Binding Udp-2F-Galactose to Alpha-1,3 Galactosyltransferase-Implications for Catalysis
Jamaluddin, H., Tumbale, P., Withers, S.G., Acharya, K.R., Brew, K.(2007) J Mol Biol 369: 1270
- PubMed: 17493636
- DOI: https://doi.org/10.1016/j.jmb.2007.04.012
- Primary Citation of Related Structures:
2JCJ, 2JCK, 2JCL, 2JCO, 2VFZ - PubMed Abstract:
Alpha-1,3 galactosyltransferase (alpha3GT) catalyzes the transfer of galactose from UDP-galactose to beta-linked galactosides with retention of its alpha configuration. Although several complexes of alpha3GT with inhibitors and substrates have been reported, no structure has been determined of a complex containing intact UDP-galactose. We describe the structure of a complex containing an inhibitory analogue of UDP-galactose, UDP-2F-galactose, in a complex with the Arg365Lys mutant of alpha3GT. The inhibitor is bound in a distorted, bent configuration and comparison with the structure of the apo form of this mutant shows that the interaction induces structural changes in the enzyme, implying a role for ground state destabilization in catalysis. In addition to a general reduction in flexibility in the enzyme indicated by a large reduction in crystallographic B-factors, two loops, one centred around Trp195 and one encompassing the C-terminal 11 residues undergo large structural changes in complexes with UDP and UDP derivatives. The distorted configuration of the bound UDP-2F-galactose in its complex is stabilized, in part, by interactions with residues that are part of or near the flexible loops. Mutagenesis and truncation studies indicate that two highly conserved basic amino acid residues in the C-terminal region, Lys359 and Arg365 are important for catalysis, probably reflecting their roles in these ligand-mediated conformational changes. A second Mn(2+) cofactor has been identified in the catalytic site of a complex of the Arg365Lys with UDP, in a location that suggests it could play a role in facilitating UDP release, consistent with kinetic studies that show alpha3GT activity depends on the binding of two manganese ions. Conformational changes in the C-terminal 11 residues require an initial reorganization of the Trp195 loop and are linked to enzyme progress through the catalytic cycle, including donor substrate distortion, cleavage of the UDP-galactose bond, galactose transfer, and UDP release.
Organizational Affiliation:
Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, UK.