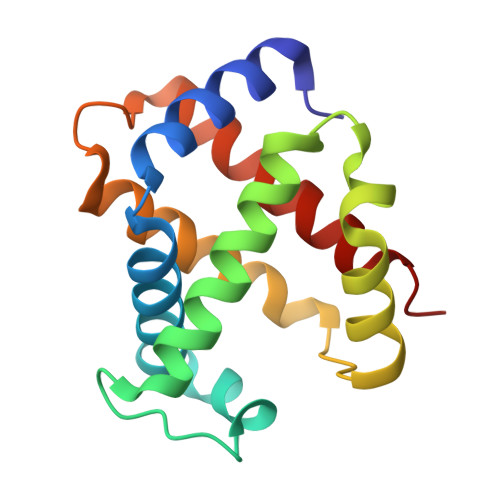
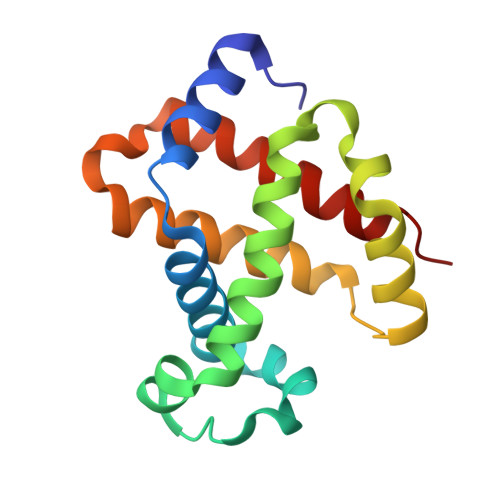
Pattern of Cavities in Globins: The Case of Human Hemoglobin.
Savino, C., Miele, A.E., Draghi, F., Johnson, K.A., Sciara, G., Brunori, M., Vallone, B.(2009) Biopolymers 91: 1097
- PubMed: 19365817
- DOI: https://doi.org/10.1002/bip.21201
- Primary Citation of Related Structures:
2W6V, 2W6W, 2W72 - PubMed Abstract:
Our aim is to shed light on the conservation of potential ligand docking sites that play an important role in ligand dynamics of globins by using the technique of filling internal cavities naturally present in hemoglobin and myoglobin with xenon atoms. In particular, we present the high resolution structures of the Xe-adduct of deoxygenated wild type human hemoglobin and a quadruple mutant (L(B10)Y and H(E7)Q in alpha and beta chains). For the sake of comparison we also determined under the same experimental conditions the xenon complex of wild type sperm whale myoglobin. The analysis revealed that the number and position of Xe binding cavities are different in the alpha and beta subunits, the latter being more similar to myoglobin. Notably, no proximal Xe docking site was detected in hemoglobin, at variance with myoglobin. The pattern of internal cavities accessibility and affinity for xenon suggests a different role for the dynamics of ligand migration in the two types of hemoglobin chains as compared to myoglobin. The number and position of hydrophobic cavities in hemoglobin are briefly discussed also in comparison with the data available for other members of the globin superfamily.
Organizational Affiliation:
University of Rome, P.le Aldo Moro 5, 00185 Rome, Italy. [email protected]