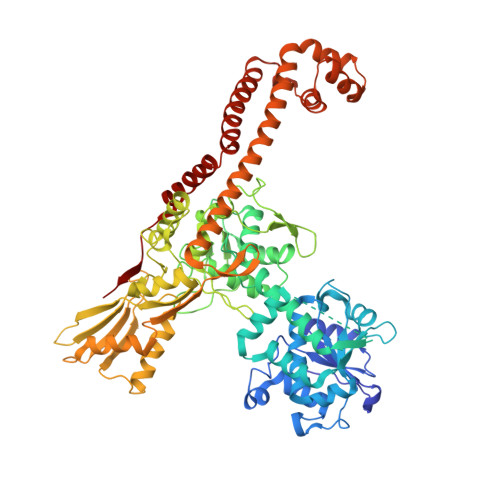
Type Iia Topoisomerase Inhibition by a New Class of Antibacterial Agents.
Bax, B.D., Chan, P.F., Eggleston, D.S., Fosberry, A., Gentry, D.R., Gorrec, F., Giordano, I., Hann, M.M., Hennessy, A., Hibbs, M., Huang, J., Jones, E., Jones, J., Brown, K.K., Lewis, C.J., May, E.W., Saunders, M.R., Singh, O., Spitzfaden, C., Shen, C., Shillings, A., Theobald, A.F., Wohlkonig, A., Pearson, N.D., Gwynn, M.N.(2010) Nature 466: 935
- PubMed: 20686482
- DOI: https://doi.org/10.1038/nature09197
- Primary Citation of Related Structures:
2XCO, 2XCQ, 2XCR, 2XCS, 2XCT - PubMed Abstract:
Despite the success of genomics in identifying new essential bacterial genes, there is a lack of sustainable leads in antibacterial drug discovery to address increasing multidrug resistance. Type IIA topoisomerases cleave and religate DNA to regulate DNA topology and are a major class of antibacterial and anticancer drug targets, yet there is no well developed structural basis for understanding drug action. Here we report the 2.1 A crystal structure of a potent, new class, broad-spectrum antibacterial agent in complex with Staphylococcus aureus DNA gyrase and DNA, showing a new mode of inhibition that circumvents fluoroquinolone resistance in this clinically important drug target. The inhibitor 'bridges' the DNA and a transient non-catalytic pocket on the two-fold axis at the GyrA dimer interface, and is close to the active sites and fluoroquinolone binding sites. In the inhibitor complex the active site seems poised to cleave the DNA, with a single metal ion observed between the TOPRIM (topoisomerase/primase) domain and the scissile phosphate. This work provides new insights into the mechanism of topoisomerase action and a platform for structure-based drug design of a new class of antibacterial agents against a clinically proven, but conformationally flexible, enzyme class.
Organizational Affiliation:
Molecular Discovery Research, GlaxoSmithKline, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire, SG1 2NY, UK. [email protected]