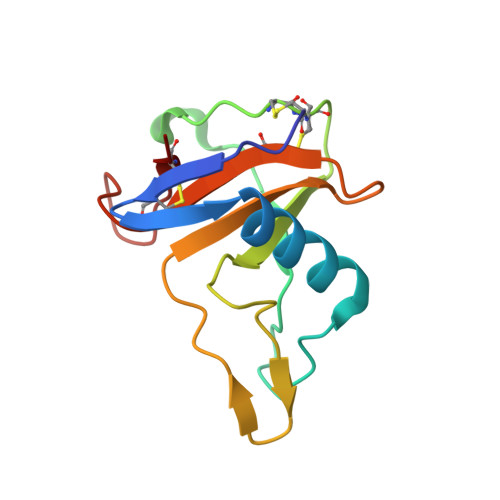
Isomerization mechanism of aspartate to isoaspartate implied by structures of Ustilago sphaerogena ribonuclease U2 complexed with adenosine 3'-monophosphate
Noguchi, S.(2010) Acta Crystallogr D Biol Crystallogr 66: 843-849
- PubMed: 20606265
- DOI: https://doi.org/10.1107/S0907444910019621
- Primary Citation of Related Structures:
3AGN, 3AGO - PubMed Abstract:
Aspartates in proteins are isomerized non-enzymatically to isoaspartate via succinimide in vitro and in vivo. In order to elucidate the mechanism of isoaspartate formation within the Asp45-Glu46 sequence of Ustilago sphaerogena ribonuclease U2 based on three-dimensional structure, crystal structures of ribonuclease U2 complexed with adenosine 3'-monophosphate have been solved at 0.96 and 0.99 A resolution. The crystal structures revealed that the C(gamma) atom of Asp45 is located just beside the main-chain N atom of Glu46 and that the conformation which is suitable for succinimide formation is stabilized by a hydrogen-bond network mediated by water molecules 190, 219 and 220. These water molecules are suggested to promote the formation of isoaspartate via succinimide: in the succinimide-formation reaction water 219 receives a proton from the N atom of Glu46 as a general base and waters 190 and 220 stabilize the tetrahedral intermediate, and in the succinimide-hydrolysis reaction water 219 provides a proton for the N atom of Glu46 as a general acid. The purine-base recognition scheme of ribonuclease U2 is also discussed.
Organizational Affiliation:
Graduate School of Pharmaceutical Sciences, The University of Tokyo, Bunkyo-ku, Tokyo, Japan. [email protected]