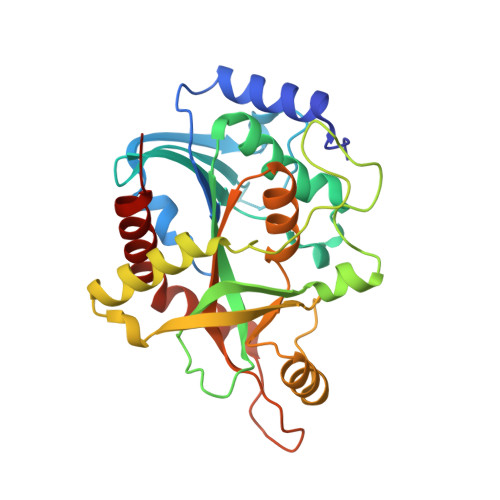
Purine nucleoside phosphorylase from Schistosoma mansoni in complex with ribose-1-phosphate.
D'Muniz Pereira, H., Oliva, G., Garratt, R.C.(2011) J Synchrotron Radiat 18: 62-65
- PubMed: 21169694
- DOI: https://doi.org/10.1107/S0909049510027718
- Primary Citation of Related Structures:
3FB1 - PubMed Abstract:
Schistosomes are blood flukes which cause schistosomiasis, a disease affecting approximately 200 million people worldwide. Along with several other important human parasites including trypanosomes and Plasmodium, schistosomes lack the de novo pathway for purine synthesis and depend exclusively on the salvage pathway for their purine requirements, making the latter an attractive target for drug development. Part of the pathway involves the conversion of inosine (or guanosine) into hypoxanthine (or guanine) together with ribose-1-phosphate (R1P) or vice versa. This inter-conversion is undertaken by the enzyme purine nucleoside phosphorylase (PNP) which has been used as the basis for the development of novel anti-malarials, conceptually validating this approach. It has been suggested that, during the reverse reaction, R1P binding to the enzyme would occur only as a consequence of conformational changes induced by hypoxanthine, thus making a binary PNP-R1P complex unlikely. Contradictory to this statement, a crystal structure of just such a binary complex involving the Schistosoma mansoni enzyme has been successfully obtained. The ligand shows an intricate hydrogen-bonding network in the phosphate and ribose binding sites and adds a further chapter to our knowledge which could be of value in the future development of selective inhibitors.
Organizational Affiliation:
Centro de Biotecnologia Molecular Estrutural, Instituto de Física de São Carlos, Universidade de São Paulo, Avenida Trabalhador São-Carlense 400, CEP 13566-590, Brazil. [email protected]