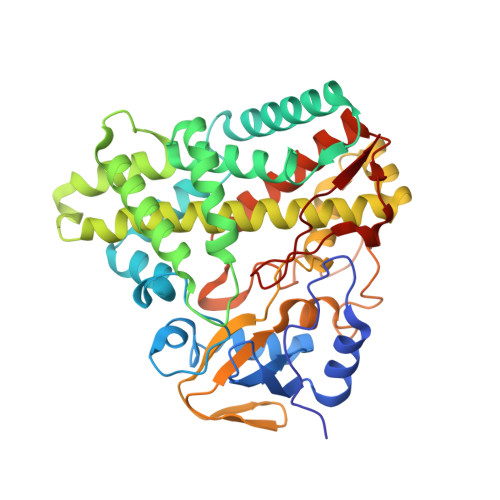
Identification and structural basis of the reaction catalyzed by CYP121, an essential cytochrome P450 in Mycobacterium tuberculosis.
Belin, P., Le Du, M.H., Fielding, A., Lequin, O., Jacquet, M., Charbonnier, J.B., Lecoq, A., Thai, R., Courcon, M., Masson, C., Dugave, C., Genet, R., Pernodet, J.L., Gondry, M.(2009) Proc Natl Acad Sci U S A 106: 7426-7431
- PubMed: 19416919
- DOI: https://doi.org/10.1073/pnas.0812191106
- Primary Citation of Related Structures:
3G5F, 3G5H - PubMed Abstract:
The gene encoding the cytochrome P450 CYP121 is essential for Mycobacterium tuberculosis. However, the CYP121 catalytic activity remains unknown. Here, we show that the cyclodipeptide cyclo(l-Tyr-l-Tyr) (cYY) binds to CYP121, and is efficiently converted into a single major product in a CYP121 activity assay containing spinach ferredoxin and ferredoxin reductase. NMR spectroscopy analysis of the reaction product shows that CYP121 catalyzes the formation of an intramolecular C-C bond between 2 tyrosyl carbon atoms of cYY resulting in a novel chemical entity. The X-ray structure of cYY-bound CYP121, solved at high resolution (1.4 A), reveals one cYY molecule with full occupancy in the large active site cavity. One cYY tyrosyl approaches the heme and establishes a specific H-bonding network with Ser-237, Gln-385, Arg-386, and 3 water molecules, including the sixth iron ligand. These observations are consistent with low temperature EPR spectra of cYY-bound CYP121 showing a change in the heme environment with the persistence of the sixth heme iron ligand. As the carbon atoms involved in the final C-C coupling are located 5.4 A apart according to the CYP121-cYY complex crystal structure, we propose that C-C coupling is concomitant with substrate tyrosyl movements. This study provides insight into the catalytic activity, mechanism, and biological function of CYP121. Also, it provides clues for rational design of putative CYP121 substrate-based antimycobacterial agents.
Organizational Affiliation:
Service d'Ingénierie Moléculaire des Protéines, Biologie Structurale et Mécanismes, Commissariat à l'Energie Atomique, Institut de Biologie et Technologies de Saclay, F-91191 Gif-sur-Yvette, France. [email protected]