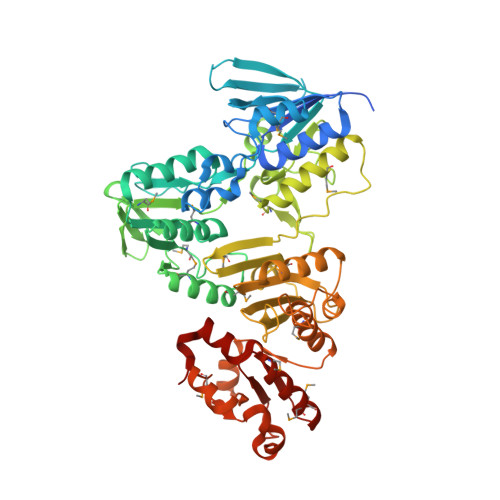
Crystal structure and catalytic properties of Bacillus anthracis CoADR-RHD: implications for flavin-linked sulfur trafficking.
Wallen, J.R., Mallett, T.C., Boles, W., Parsonage, D., Furdui, C.M., Karplus, P.A., Claiborne, A.(2009) Biochemistry 48: 9650-9667
- PubMed: 19725515
- DOI: https://doi.org/10.1021/bi900887k
- Primary Citation of Related Structures:
3ICR, 3ICS, 3ICT - PubMed Abstract:
Rhodanese homology domains (RHDs) play important roles in sulfur trafficking mechanisms essential to the biosynthesis of sulfur-containing cofactors and nucleosides. We have now determined the crystal structure at 2.10 A resolution for the Bacillus anthracis coenzyme A-disulfide reductase isoform (BaCoADR-RHD) containing a C-terminal RHD domain; this is the first structural representative of the multidomain proteins class of the rhodanese superfamily. The catalytic Cys44 of the CoADR module is separated by 25 A from the active-site Cys514' of the RHD domain from the complementary subunit. In stark contrast to the B. anthracis CoADR [Wallen, J. R., Paige, C., Mallett, T. C., Karplus, P. A., and Claiborne, A. (2008) Biochemistry 47, 5182-5193], the BaCoADR-RHD isoform does not catalyze the reduction of coenzyme A-disulfide, although both enzymes conserve the Cys-SSCoA redox center. NADH titrations have been combined with a synchrotron reduction protocol for examination of the structural and redox behavior of the Cys44-SSCoA center. The synchrotron-reduced (Cys44 + CoASH) structure reveals ordered binding for the adenosine 3'-phosphate 5'-pyrophosphate moiety of CoASH, but the absence of density for the pantetheine arm indicates that it is flexible within the reduced active site. Steady-state kinetic analyses with the alternate disulfide substrates methyl methanethiolsulfonate (MMTS) and 5,5'-dithiobis(2-nitrobenzoate) (DTNB), including the appropriate Cys --> Ser mutants, demonstrate that MMTS reduction occurs within the CoADR active site. NADH-dependent DTNB reduction, on the other hand, requires communication between Cys44 and Cys514', and we propose that reduction of the Cys44-SSCoA disulfide promotes the transfer of reducing equivalents to the RHD, with the swinging pantetheine arm serving as a ca. 20 A bridge.
Organizational Affiliation:
Department of Internal Medicine, Wake Forest University Schoolof Medicine, Center for Structural Biology, Winston-Salem, North Carolina 27157, USA.