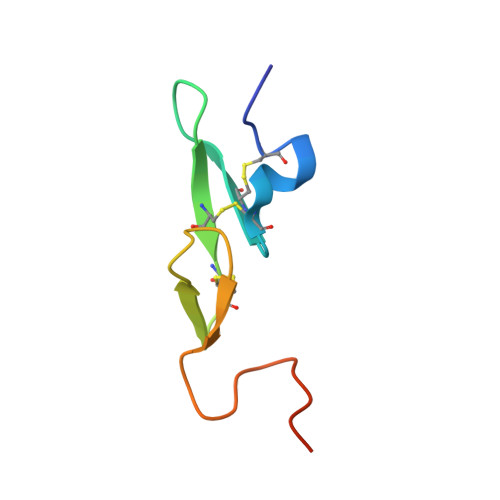
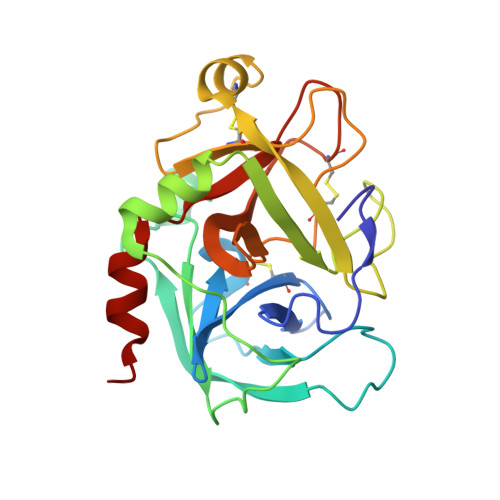
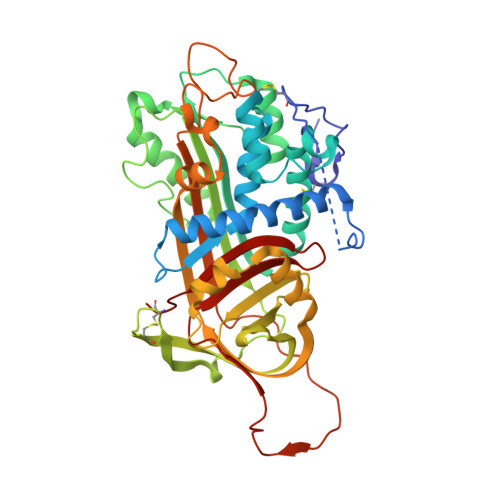
Molecular basis of factor IXa recognition by heparin-activated antithrombin revealed by a 1.7-A structure of the ternary complex.
Johnson, D.J., Langdown, J., Huntington, J.A.(2010) Proc Natl Acad Sci U S A 107: 645-650
- PubMed: 20080729
- DOI: https://doi.org/10.1073/pnas.0910144107
- Primary Citation of Related Structures:
3KCG - PubMed Abstract:
Factor (f) IXa is a critical enzyme for the formation of stable blood clots, and its deficiency results in hemophilia. The enzyme functions at the confluence of the intrinsic and extrinsic pathways by binding to fVIIIa and rapidly generating fXa. In spite of its importance, little is known about how fIXa recognizes its cofactor, its substrate, or its only known inhibitor, antithrombin (AT). However, it is clear that fIXa requires extensive exosite interactions to present substrates for efficient cleavage. Here we describe the 1.7-A crystal structure of fIXa in its recognition (Michaelis) complex with heparin-activated AT. It represents the highest resolution structure of both proteins and allows us to address several outstanding issues. The structure reveals why the heparin-induced conformational change in AT is required to permit simultaneous active-site and exosite interactions with fIXa and the nature of these interactions. The reactive center loop of AT has evolved to specifically inhibit fIXa, with a P2 Gly so as not to clash with Tyr99 on fIXa, a P4 Ile to fit snugly into the S4 pocket, and a C-terminal extension to exploit a unique wall-like feature of the active-site cleft. Arg150 is at the center of the exosite interface, interacting with AT residues on beta-sheet C. A surprising crystal contact is observed between the heparin pentasaccharide and fIXa, revealing a plausible mode of binding that would allow longer heparin chains to bridge the complex.
Organizational Affiliation:
Department of Haematology, Cambridge Institute for Medical Research, University of Cambridge, Cambridge CB2 0XY, United Kingdom.