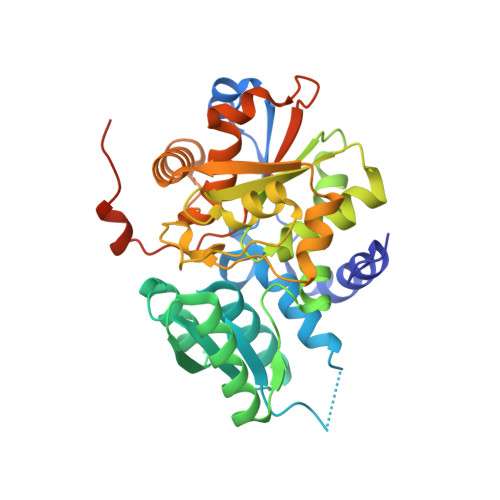
The structure of mammalian serine racemase: evidence for conformational changes upon inhibitor binding.
Smith, M.A., Mack, V., Ebneth, A., Moraes, I., Felicetti, B., Wood, M., Schonfeld, D., Mather, O., Cesura, A., Barker, J.(2010) J Biol Chem 285: 12873-12881
- PubMed: 20106978
- DOI: https://doi.org/10.1074/jbc.M109.050062
- Primary Citation of Related Structures:
3HMK, 3L6B, 3L6C, 3L6R - PubMed Abstract:
Serine racemase is responsible for the synthesis of D-serine, an endogenous co-agonist for N-methyl-D-aspartate receptor-type glutamate receptors (NMDARs). This pyridoxal 5'-phosphate-dependent enzyme is involved both in the reversible conversion of L- to D-serine and serine catabolism by alpha,beta-elimination of water, thereby regulating D-serine levels. Because D-serine affects NMDAR signaling throughout the brain, serine racemase is a promising target for the treatment of disorders related to NMDAR dysfunction. To provide a molecular basis for rational drug design the x-ray crystal structures of human and rat serine racemase were determined at 1.5- and 2.1-A resolution, respectively, and in the presence and absence of the orthosteric inhibitor malonate. The structures revealed a fold typical of beta-family pyridoxal 5'-phosphate enzymes, with both a large domain and a flexible small domain associated into a symmetric dimer, and indicated a ligand-induced rearrangement of the small domain that organizes the active site for specific turnover of the substrate.
Organizational Affiliation:
Department of Structural Biology and Biology, Evotec, 114 Milton Park, Abingdon, Oxon OX14 4SA, United Kingdom. [email protected]