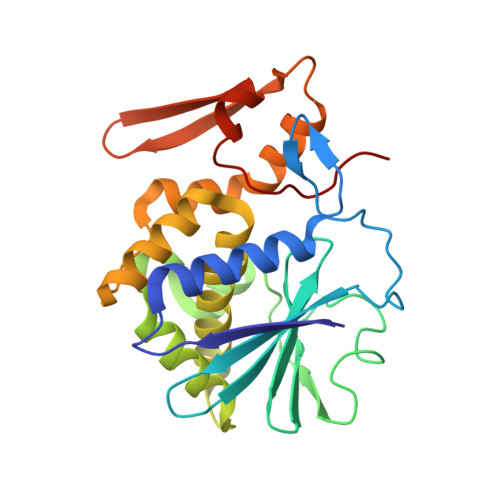
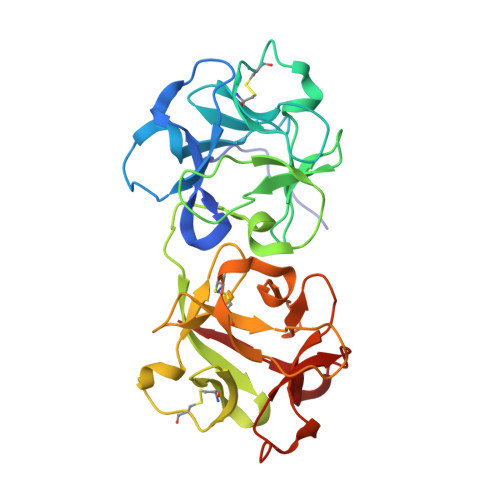
Binding of the plant hormone kinetin in the active site of Mistletoe Lectin I from Viscum album.
Malecki, P.H., Rypniewski, W., Szymanski, M., Barciszewski, J., Meyer, A.(2012) Biochim Biophys Acta 1824: 334-338
- PubMed: 22064121
- DOI: https://doi.org/10.1016/j.bbapap.2011.10.013
- Primary Citation of Related Structures:
3O5W - PubMed Abstract:
The crystal structure of the ribosome inhibiting protein Mistletoe Lectin I (ML-I) derived from the European mistletoe, Viscum album, in complex with kinetin has been refined at 2.7Å resolution. Suitably large crystals of ML-I were obtained applying the counter diffusion method using the Gel Tube R Crystallization Kit (GT-R) on board the Russian Service Module on the international space station ISS within the GCF mission No. 6, arranged by the Japanese aerospace exploration agency (JAXA). Hexagonal bi-pyramidal crystals were grown during three months under microgravity. Before data collection the crystals were soaked in a saturated solution of kinetin and diffraction data to 2.7Å were collected using synchrotron radiation and cryogenic techniques. The atomic model was refined and revealed a single kinetin molecule in the ribosome inactivation site of ML-I. The complex demonstrates the feasibility of mistletoe to bind plant hormones out of the host regulation system as part of a self protection mechanism.
Organizational Affiliation:
Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan, Poland.