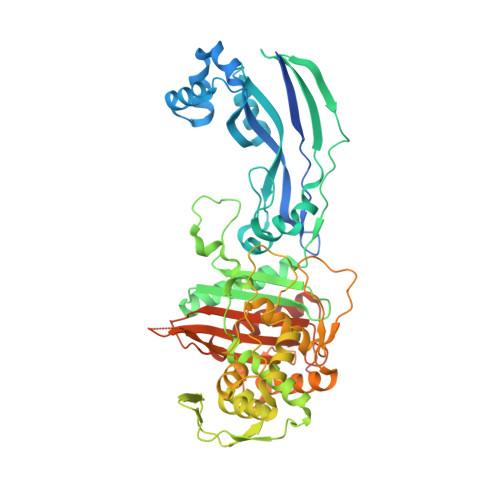
Crystal structures of penicillin-binding protein 3 from Pseudomonas aeruginosa: comparison of native and antibiotic-bound forms
Sainsbury, S., Bird, L., Rao, V., Shepherd, S.M., Stuart, D.I., Hunter, W.N., Owens, R.J., Ren, J.(2011) J Mol Biol 405: 173-184
- PubMed: 20974151
- DOI: https://doi.org/10.1016/j.jmb.2010.10.024
- Primary Citation of Related Structures:
3OC2, 3OCL, 3OCN - PubMed Abstract:
We report the first crystal structures of a penicillin-binding protein (PBP), PBP3, from Pseudomonas aeruginosa in native form and covalently linked to two important β-lactam antibiotics, carbenicillin and ceftazidime. Overall, the structures of apo and acyl complexes are very similar; however, variations in the orientation of the amino-terminal membrane-proximal domain relative to that of the carboxy-terminal transpeptidase domain indicate interdomain flexibility. Binding of either carbenicillin or ceftazidime to purified PBP3 increases the thermostability of the enzyme significantly and is associated with local conformational changes, which lead to a narrowing of the substrate-binding cleft. The orientations of the two β-lactams in the active site and the key interactions formed between the ligands and PBP3 are similar despite differences in the two drugs, indicating a degree of flexibility in the binding site. The conserved binding mode of β-lactam-based inhibitors appears to extend to other PBPs, as suggested by a comparison of the PBP3/ceftazidime complex and the Escherichia coli PBP1b/ceftoxamine complex. Since P. aeruginosa is an important human pathogen, the structural data reveal the mode of action of the frontline antibiotic ceftazidime at the molecular level. Improved drugs to combat infections by P. aeruginosa and related Gram-negative bacteria are sought and our study provides templates to assist that process and allows us to discuss new ways of inhibiting PBPs.
Organizational Affiliation:
Henry Wellcome Building for Genomic Medicine, University of Oxford, Oxford OX3 7BN, UK.