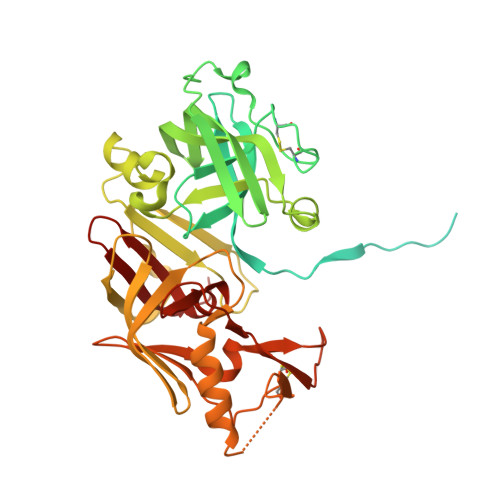
Structural insights into the activation and inhibition of histo-aspartic protease from Plasmodium falciparum.
Bhaumik, P., Xiao, H., Hidaka, K., Gustchina, A., Kiso, Y., Yada, R.Y., Wlodawer, A.(2011) Biochemistry 50: 8862-8879
- PubMed: 21928835
- DOI: https://doi.org/10.1021/bi201118z
- Primary Citation of Related Structures:
3QVC, 3QVI - PubMed Abstract:
Histo-aspartic protease (HAP) from Plasmodium falciparum is a promising target for the development of novel antimalarial drugs. The sequence of HAP is highly similar to those of pepsin-like aspartic proteases, but one of the two catalytic aspartates, Asp32, is replaced with histidine. Crystal structures of the truncated zymogen of HAP and of the complex of the mature enzyme with inhibitor KNI-10395 have been determined at 2.1 and 2.5 Å resolution, respectively. As in other proplasmepsins, the propeptide of the zymogen interacts with the C-terminal domain of the enzyme, forcing the N- and C-terminal domains apart, thereby separating His32 and Asp215 and preventing formation of the mature active site. In the inhibitor complex, the enzyme forms a tight domain-swapped dimer, not previously seen in any aspartic proteases. The inhibitor is found in an unprecedented conformation resembling the letter U, stabilized by two intramolecular hydrogen bonds. Surprisingly, the location and conformation of the inhibitor are similar to those of the fragment of helix 2 comprising residues 34p-38p in the prosegments of the zymogens of gastric aspartic proteases; a corresponding helix assumes a vastly different orientation in proplasmepsins. Each inhibitor molecule is in contact with two molecules of HAP, interacting with the carboxylate group of the catalytic Asp215 of one HAP protomer through a water molecule, while also making a direct hydrogen bond to Glu278A' of the other protomer. A comparison of the shifts in the positions of the catalytic residues in the inhibitor complex presented here with those published previously gives further hints regarding the enzymatic mechanism of HAP.
Organizational Affiliation:
Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute, Frederick, Maryland 21702, USA.