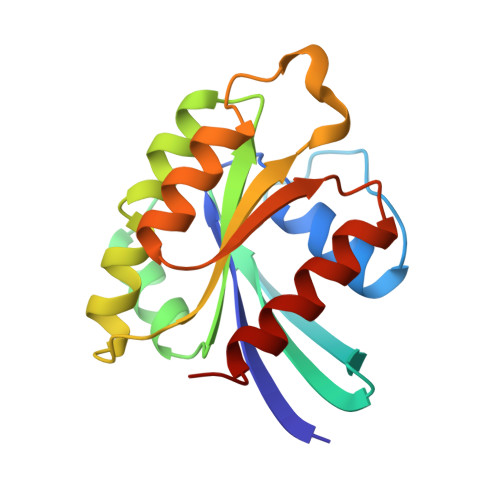
Structure of the small G protein Rap2 in a non-catalytic complex with GTP.
Menetrey, J., Cherfils, J.(1999) Proteins 37: 465-473
- PubMed: 10591105
- DOI: https://doi.org/10.1002/(sici)1097-0134(19991115)37:3<465::aid-prot13>3.0.co;2-o
- Primary Citation of Related Structures:
3RAP - PubMed Abstract:
We report a novel crystal form of the small G protein Rap2A in complex with GTP which has no GTPase activity in the crystal. The asymmetric unit contains two complexes which show that a conserved switch I residue, Tyr 32, contributes an extra hydrogen bond to the gamma-phosphate of GTP as compared to related structures with GTP analogs. Since GTP is not hydrolyzed in the crystal, this interaction is unlikely to contribute to the intrinsic GTPase activity. The comparison of other G protein structures to the Rap2-GTP complex suggests that an equivalent interaction is likely to exist in their GTP form, whether unbound or bound to an effector. This interaction has to be released to allow the GAP-activated GTPase, and presumably the intrinsic GTPase activity as well. We also discuss the definition of the flexible regions and their hinges in the light of this structure and the expanding database of G protein structures. We propose that the switch I and switch II undergo either partial or complete disorder-to-order transitions according to their cellular status, thus defining a complex energy landscape comprising more than two conformational states. We observe in addition that the region connecting the switch I and switch II is flexible in Rap2 and other G proteins. This region may be important for protein-protein interactions and possibly behave as a conformational lever arm, as characterized for Arf. Taken together, these observations suggest that the structural mechanisms of small G proteins are significantly driven by entropy-based free energy changes.
Organizational Affiliation:
Laboratoire d'Enzymologie et Biochimie Structurales, Centre National de la Recherche Scientifique, Gif sur Yvette, France.