Unique Interplay between Sugar and Lipid in Determining the Antigenic Potency of Bacterial Antigens for NKT Cells.
Girardi, E., Yu, E.D., Li, Y., Tarumoto, N., Pei, B., Wang, J., Illarionov, P., Kinjo, Y., Kronenberg, M., Zajonc, D.M.(2011) PLoS Biol 9: e1001189-e1001189
- PubMed: 22069376
- DOI: https://doi.org/10.1371/journal.pbio.1001189
- Primary Citation of Related Structures:
3TA3 - PubMed Abstract:
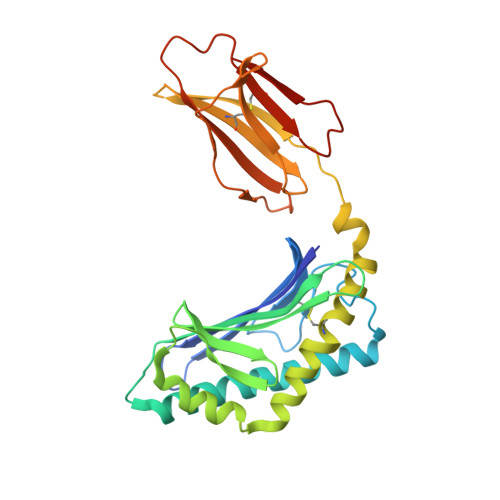
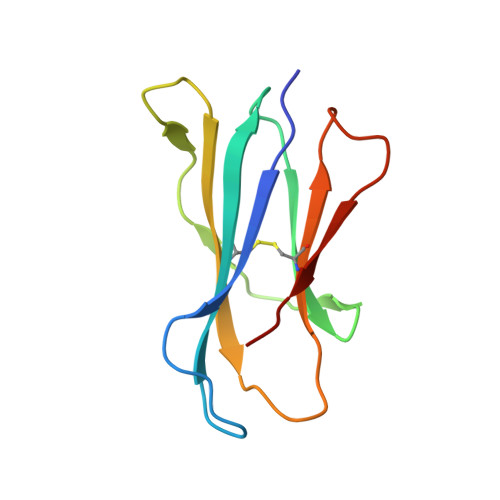
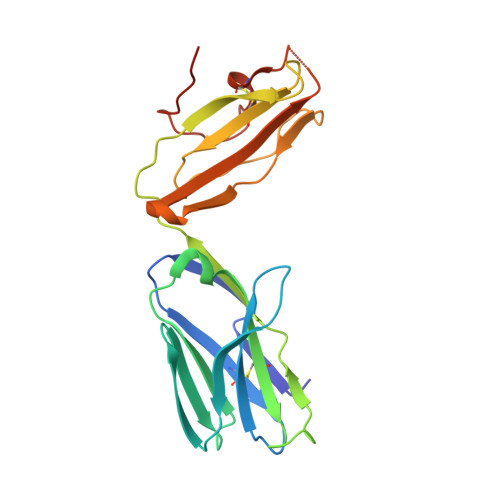
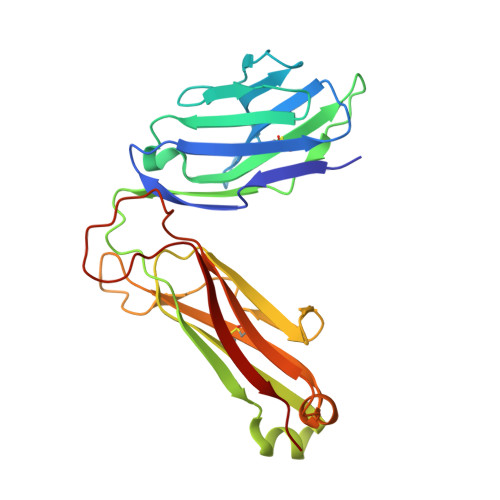
Invariant natural killer T (iNKT) cells are an evolutionary conserved T cell population characterized by features of both the innate and adaptive immune response. Studies have shown that iNKT cells are required for protective responses to Gram-positive pathogens such as Streptococcus pneumoniae, and that these cells recognize bacterial diacylglycerol antigens presented by CD1d, a non-classical antigen-presenting molecule. The combination of a lipid backbone containing an unusual fatty acid, vaccenic acid, as well as a glucose sugar that is weaker or not stimulatory when linked to other lipids, is required for iNKT cell stimulation by these antigens. Here we have carried out structural and biophysical studies that illuminate the reasons for the stringent requirement for this unique combination. The data indicate that vaccenic acid bound to the CD1d groove orients the protruding glucose sugar for TCR recognition, and it allows for an additional hydrogen bond of the glucose with CD1d when in complex with the TCR. Furthermore, TCR binding causes an induced fit in both the sugar and CD1d, and we have identified the CD1d amino acids important for iNKT TCR recognition and the stability of the ternary complex. The studies show also how hydrogen bonds formed by the glucose sugar can account for the distinct binding kinetics of the TCR for this CD1d-glycolipid complex. Therefore, our studies illuminate the mechanism of glycolipid recognition for antigens from important pathogens.
Organizational Affiliation:
Division of Cell Biology, La Jolla Institute for Allergy & Immunology, La Jolla, California, United States of America.