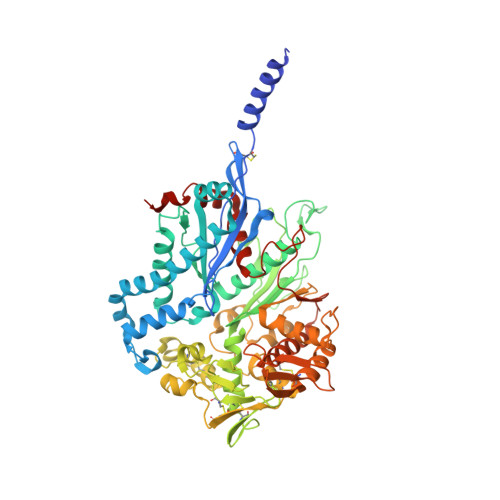
Structural Insight Into the Activation Mechanism of Toxoplasma Gondii Nucleoside Triphosphate Diphosphohydrolases by Disulfide Reduction.
Krug, U., Zebisch, M., Krauss, M., Straeter, N.(2012) J Biol Chem 287: 3051
- PubMed: 22130673
- DOI: https://doi.org/10.1074/jbc.M111.294348
- Primary Citation of Related Structures:
4A57, 4A59, 4A5A, 4A5B - PubMed Abstract:
The intracellular parasite Toxoplasma gondii produces two nucleoside triphosphate diphosphohydrolases (NTPDase1 and -3). These tetrameric, cysteine-rich enzymes require activation by reductive cleavage of a hitherto unknown disulfide bond. Despite a 97% sequence identity, both isozymes differ largely in their ability to hydrolyze ATP and ADP. Here, we present crystal structures of inactive NTPDase3 as an apo form and in complex with the product AMP to resolutions of 2.0 and 2.2 Å, respectively. We find that the enzyme is present in an open conformation that precludes productive substrate binding and catalysis. The cysteine bridge 258-268 is identified to be responsible for locking of activity. Crystal structures of constitutively active variants of NTPDase1 and -3 generated by mutation of Cys(258)-Cys(268) show that opening of the regulatory cysteine bridge induces a pronounced contraction of the whole tetramer. This is accompanied by a 12° domain closure motion resulting in the correct arrangement of all active site residues. A complex structure of activated NTPDase3 with a non-hydrolyzable ATP analog and the cofactor Mg(2+) to a resolution of 2.85 Å indicates that catalytic differences between the NTPDases are primarily dictated by differences in positioning of the adenine base caused by substitution of Arg(492) and Glu(493) in NTPDase1 by glycines in NTPDase3.
Organizational Affiliation:
Institute of Bioanalytical Chemistry, Center for Biotechnology and Biomedicine, University of Leipzig, D-04103 Leipzig, Germany.