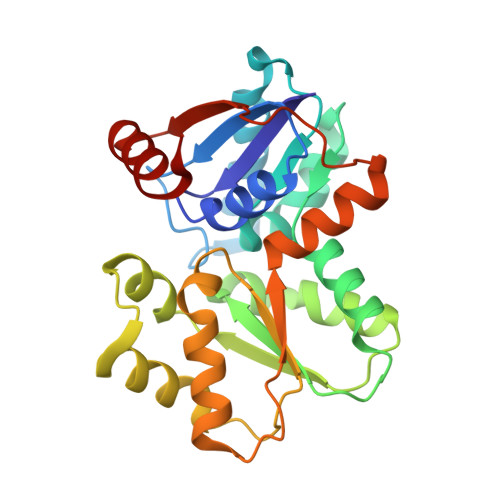
Design of Inhibitors of Helicobacter Pylori Glutamate Racemase as Selective Antibacterial Agents: Incorporation of Imidazoles Onto a Core Pyrazolopyrimidinedione Scaffold to Improve Bioavailabilty.
Basarab, G.S., Hill, P., Eyermann, C.J., Gowravaram, M., Kack, H., Osimoni, E.(2012) Bioorg Med Chem Lett 22: 5600
- PubMed: 22877632
- DOI: https://doi.org/10.1016/j.bmcl.2012.07.004
- Primary Citation of Related Structures:
4B1F - PubMed Abstract:
Structure-activity relationships are presented around a series of pyrazolopyrimidinediones that inhibit the growth of Helicobacter pylori by targeting glutamate racemase, an enzyme that provides d-glutamate for the construction of N-acetylglucosamine-N-acetylmuramic acid peptidoglycan subunits assimilated into the bacterial cell wall. Substituents on the inhibitor scaffold were varied to optimize target potency, antibacterial activity and in vivo pharmacokinetic stability. By incorporating an imidazole ring at the 7-position of scaffold, high target potency was achieved due to a hydrogen bonding network that occurs between the 3-position nitrogen atom, a bridging water molecule and the side chains Ser152 and Trp244 of the enzyme. The lipophilicity of the scaffold series proved important for expression of antibacterial activity. Clearances in vitro and in vivo were monitored to identify compounds with improved plasma stability. The basicity of the imidazole may contribute to increased aqueous solubility at lower pH allowing for improved oral bioavailability.
Organizational Affiliation:
Infection Innovative Medicines, AstraZeneca R&D Boston, 35 Gatehouse Drive, Waltham, MA 02451, USA. [email protected]