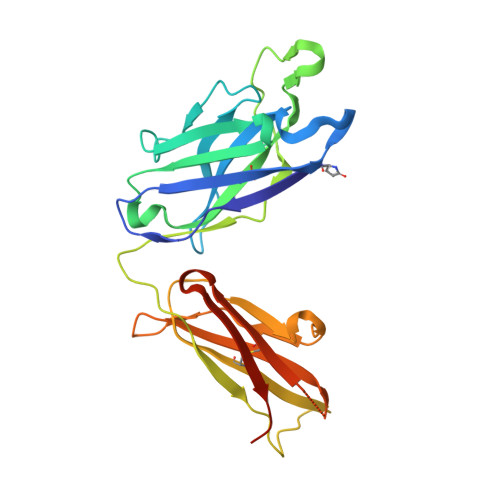
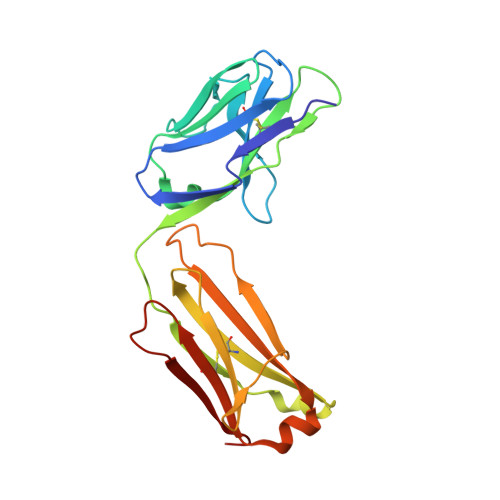
Cross-neutralization of influenza A viruses mediated by a single antibody loop.
Ekiert, D.C., Kashyap, A.K., Steel, J., Rubrum, A., Bhabha, G., Khayat, R., Lee, J.H., Dillon, M.A., O'Neil, R.E., Faynboym, A.M., Horowitz, M., Horowitz, L., Ward, A.B., Palese, P., Webby, R., Lerner, R.A., Bhatt, R.R., Wilson, I.A.(2012) Nature 489: 526-532
- PubMed: 22982990
- DOI: https://doi.org/10.1038/nature11414
- Primary Citation of Related Structures:
4FNK, 4FNL, 4FP8, 4FQR - PubMed Abstract:
Immune recognition of protein antigens relies on the combined interaction of multiple antibody loops, which provide a fairly large footprint and constrain the size and shape of protein surfaces that can be targeted. Single protein loops can mediate extremely high-affinity binding, but it is unclear whether such a mechanism is available to antibodies. Here we report the isolation and characterization of an antibody called C05, which neutralizes strains from multiple subtypes of influenza A virus, including H1, H2 and H3. X-ray and electron microscopy structures show that C05 recognizes conserved elements of the receptor-binding site on the haemagglutinin surface glycoprotein. Recognition of the haemagglutinin receptor-binding site is dominated by a single heavy-chain complementarity-determining region 3 loop, with minor contacts from heavy-chain complementarity-determining region 1, and is sufficient to achieve nanomolar binding with a minimal footprint. Thus, binding predominantly with a single loop can allow antibodies to target small, conserved functional sites on otherwise hypervariable antigens.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, USA.