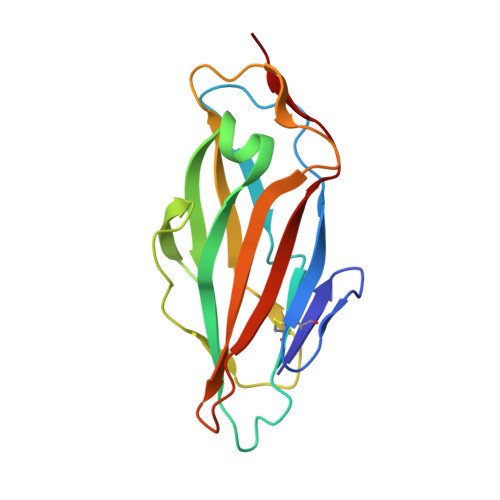
Study of the Structural and Dynamic Effects in the FimH Adhesin upon alpha-d-Heptyl Mannose Binding.
Vanwetswinkel, S., Volkov, A.N., Sterckx, Y.G., Garcia-Pino, A., Buts, L., Vranken, W.F., Bouckaert, J., Roy, R., Wyns, L., van Nuland, N.A.(2014) J Med Chem 57: 1416-1427
- PubMed: 24476493
- DOI: https://doi.org/10.1021/jm401666c
- Primary Citation of Related Structures:
3ZPD, 4LOV - PubMed Abstract:
Uropathogenic Escherichia coli cause urinary tract infections by adhering to mannosylated receptors on the human urothelium via the carbohydrate-binding domain of the FimH adhesin (FimHL). Numerous α-d-mannopyranosides, including α-d-heptyl mannose (HM), inhibit this process by interacting with FimHL. To establish the molecular basis of the high-affinity HM binding, we solved the solution structure of the apo form and the crystal structure of the FimHL-HM complex. NMR relaxation analysis revealed that protein dynamics were not affected by the sugar binding, yet HM addition promoted protein dimerization, which was further confirmed by small-angle X-ray scattering. Finally, to address the role of Y48, part of the "tyrosine gate" believed to govern the affinity and specificity of mannoside binding, we characterized the FimHL Y48A mutant, whose conformational, dynamical, and HM binding properties were found to be very similar to those of the wild-type protein.
Organizational Affiliation:
Jean Jeener NMR Centre, Structural Biology Brussels, Vrije Universiteit Brussel , Pleinlaan 2, 1050 Brussels, Belgium.