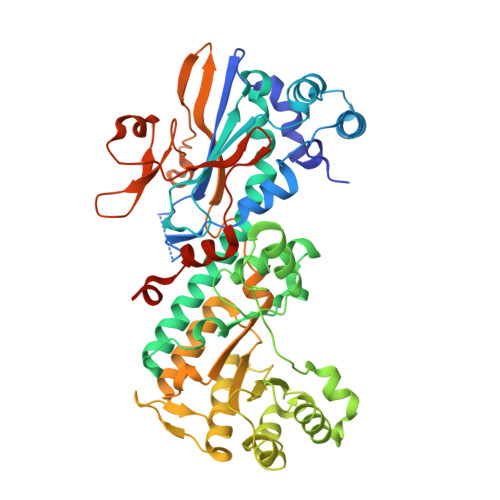
Discovery of potent and efficacious cyanoguanidine-containing nicotinamide phosphoribosyltransferase (Nampt) inhibitors.
Zheng, X., Baumeister, T., Buckmelter, A.J., Caligiuri, M., Clodfelter, K.H., Han, B., Ho, Y.C., Kley, N., Lin, J., Reynolds, D.J., Sharma, G., Smith, C.C., Wang, Z., Dragovich, P.S., Oh, A., Wang, W., Zak, M., Wang, Y., Yuen, P.W., Bair, K.W.(2014) Bioorg Med Chem Lett 24: 337-343
- PubMed: 24279990
- DOI: https://doi.org/10.1016/j.bmcl.2013.11.006
- Primary Citation of Related Structures:
4LTS, 4LWW - PubMed Abstract:
A co-crystal structure of amide-containing compound (4) in complex with the nicotinamide phosphoribosyltransferase (Nampt) protein and molecular modeling were utilized to design and discover a potent novel cyanoguanidine-containing inhibitor bearing a sulfone moiety (5, Nampt Biochemical IC50=2.5nM, A2780 cell proliferation IC50=9.7nM). Further SAR exploration identified several additional cyanoguanidine-containing compounds with high potency and good microsomal stability. Among these, compound 15 was selected for in vivo profiling and demonstrated good oral exposure in mice. It also exhibited excellent in vivo antitumor efficacy when dosed orally in an A2780 ovarian tumor xenograft model. The co-crystal structure of this compound in complex with the NAMPT protein was also determined.
Organizational Affiliation:
Forma Therapeutics, Inc., 500 Arsenal Street, Watertown, MA 02472, USA. Electronic address: [email protected].