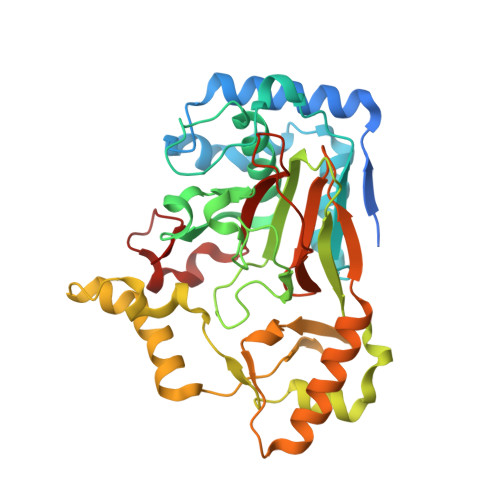
Biosynthesis of streptolidine involved two unexpected intermediates produced by a dihydroxylase and a cyclase through unusual mechanisms.
Chang, C.Y., Lyu, S.Y., Liu, Y.C., Hsu, N.S., Wu, C.C., Tang, C.F., Lin, K.H., Ho, J.Y., Wu, C.J., Tsai, M.D., Li, T.L.(2014) Angew Chem Int Ed Engl 53: 1943-1948
- PubMed: 24505011
- DOI: https://doi.org/10.1002/anie.201307989
- Primary Citation of Related Structures:
4M23, 4M25, 4M26, 4M27, 4M2C, 4M2E, 4M2F, 4M2G, 4M2I, 4M2J, 4M2K, 4M2M, 4NE0 - PubMed Abstract:
Streptothricin-F (STT-F), one of the early-discovered antibiotics, consists of three components, a β-lysine homopolymer, an aminosugar D-gulosamine, and an unusual bicyclic streptolidine. The biosynthesis of streptolidine is a long-lasting but unresolved puzzle. Herein, a combination of genetic/biochemical/structural approaches was used to unravel this problem. The STT gene cluster was first sequenced from a Streptomyces variant BCRC 12163, wherein two gene products OrfP and OrfR were characterized in vitro to be a dihydroxylase and a cyclase, respectively. Thirteen high-resolution crystal structures for both enzymes in different reaction intermediate states were snapshotted to help elucidate their catalytic mechanisms. OrfP catalyzes an Fe(II) -dependent double hydroxylation reaction converting L-Arg into (3R,4R)-(OH)2 -L-Arg via (3S)-OH-L-Arg, while OrfR catalyzes an unusual PLP-dependent elimination/addition reaction cyclizing (3R,4R)-(OH)2 -L-Arg to the six-membered (4R)-OH-capreomycidine. The biosynthetic mystery finally comes to light as the latter product was incorporation into STT-F by a feeding experiment.
Organizational Affiliation:
Genomics Research Center, Academia Sinica, 128 Academia Road, Section 2, Nankang, Taipei 115 (Taiwan).