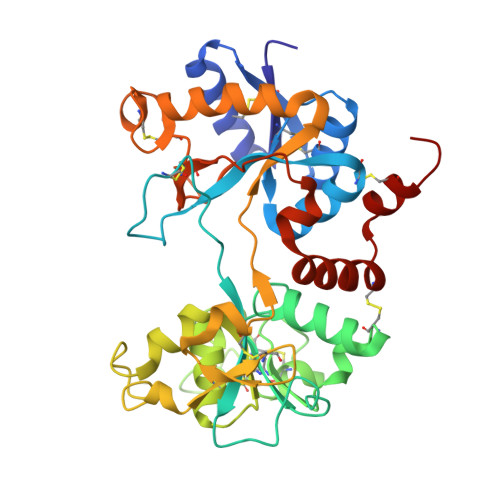
Structure of the iron-free true C-terminal half of bovine lactoferrin produced by tryptic digestion and its functional significance in the gut.
Rastogi, N., Singh, A., Pandey, S.N., Sinha, M., Bhushan, A., Kaur, P., Sharma, S., Singh, T.P.(2014) FEBS J 281: 2871-2882
- PubMed: 24798798
- DOI: https://doi.org/10.1111/febs.12827
- Primary Citation of Related Structures:
4OQO - PubMed Abstract:
Bovine lactoferrin, a 76-kDa glycoprotein (Ala1-Arg689) consists of two similar N- and C-terminal molecular halves with the ability to bind two Fe(3+) ions. The N-terminal half, designated as the N-lobe (Ala1-Arg341) and the C-terminal half designated as the C-lobe (Tyr342-Arg689) have similar iron-binding properties, but the resistant C-lobe prolongs the physiological role of bovine lactoferrin in the digestive tract. Here, we report the crystal structure of true C-lobe, which was produced by limited proteolysis of bovine lactoferrin using trypsin. In the first proteolysis step, two fragments of 21 kDa (Glu86-Lys282) and 45 kDa (Ser283-Arg689) were generated because two lysine residues, Lys85 and Lys282, in the structure of iron-saturated bovine lactoferrin were fully exposed. The 45-kDa fragment was further digested at the newly exposed side chain of Arg341, generating a 38-kDa perfect C-lobe (Tyr342-Arg689). By contrast, the apo-lactoferrin was cut by trypsin only at Arg341, which was exposed in the structure of apo-lactoferrin, whereas the other two sites with Lys85 and Lys282 are inaccessible. The purified iron-saturated C-lobe was crystallized at pH 4.0. The structure was determined by the molecular replacement method using coordinates of the C-terminal half (Arg342-Arg689) of intact camel apo-lactoferrin. The structure determination revealed that the iron atom was absent and the iron-binding cleft was found in a wide-open conformation, whereas in the previously determined structure of iron-saturated C-lobe of bovine lactoferrin, the iron atom was present and the iron-binding site was in the closed confirmation.
Organizational Affiliation:
Department of Biophysics, All India Institute of Medical Sciences, New Delhi, India.