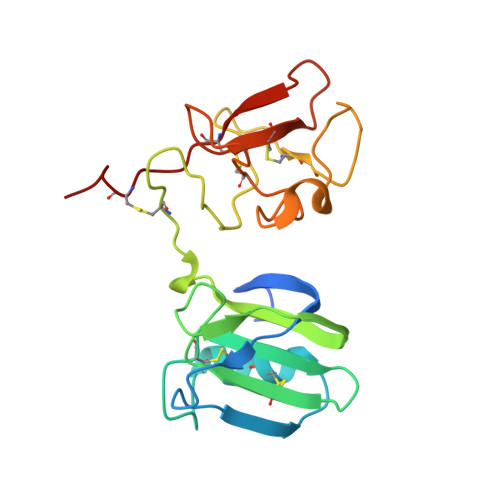
Exploring the chemical space of the lysine-binding pocket of the first kringle domain of hepatocyte growth factor/scatter factor (HGF/SF) yields a new class of inhibitors of HGF/SF-MET binding.
Sigurdardottir, A.G., Winter, A., Sobkowicz, A., Fragai, M., Chirgadze, D., Ascher, D.B., Blundell, T.L., Gherardi, E.(2015) Chem Sci 6: 6147-6157
- PubMed: 30090230
- DOI: https://doi.org/10.1039/c5sc02155c
- Primary Citation of Related Structures:
5COE, 5CP9, 5CS1, 5CS3, 5CS5, 5CS9, 5CSQ, 5CT1, 5CT2, 5CT3 - PubMed Abstract:
The growth/motility factor hepatocyte growth factor/scatter factor (HGF/SF) and its receptor, the tyrosine kinase MET, constitute a signalling system essential for embryogenesis and for tissue/organ regeneration in post-natal life. HGF/SF-MET signalling, however, also plays a key role in the onset of metastasis of a large number of human tumours. Both HGF/SF and MET are high molecular weight proteins that bury an extensive interface upon complex formation and thus constitute a challenging target for the development of low molecular weight inhibitors. Here we have used surface plasmon resonance (SPR), nuclear magnetic resonance (NMR) and X-ray crystallography to screen a diverse fragment library of 1338 members as well as a range of piperazine-like compounds. Several small molecules were found to bind in the lysine-binding pocket of the kringle 1 domain of HGF/SF and its truncated splice variant NK1. We have defined the binding mode of these compounds, explored their biological activity and we show that selected fragments inhibit MET downstream signalling. Thus we demonstrate that targeting the lysine-binding pocket of NK1 is an effective strategy to generate MET receptor antagonists and we offer proof of concept that the HGF/SF-MET interface may be successfully targeted with small molecules. These studies have broad implications for the development of HGF/SF-MET therapeutics and cancer treatment.
Organizational Affiliation:
Department of Biochemistry , University of Cambridge , 80 Tennis Court Road , Cambridge , CB2 1GA , UK . Email: [email protected] ; Email: [email protected].