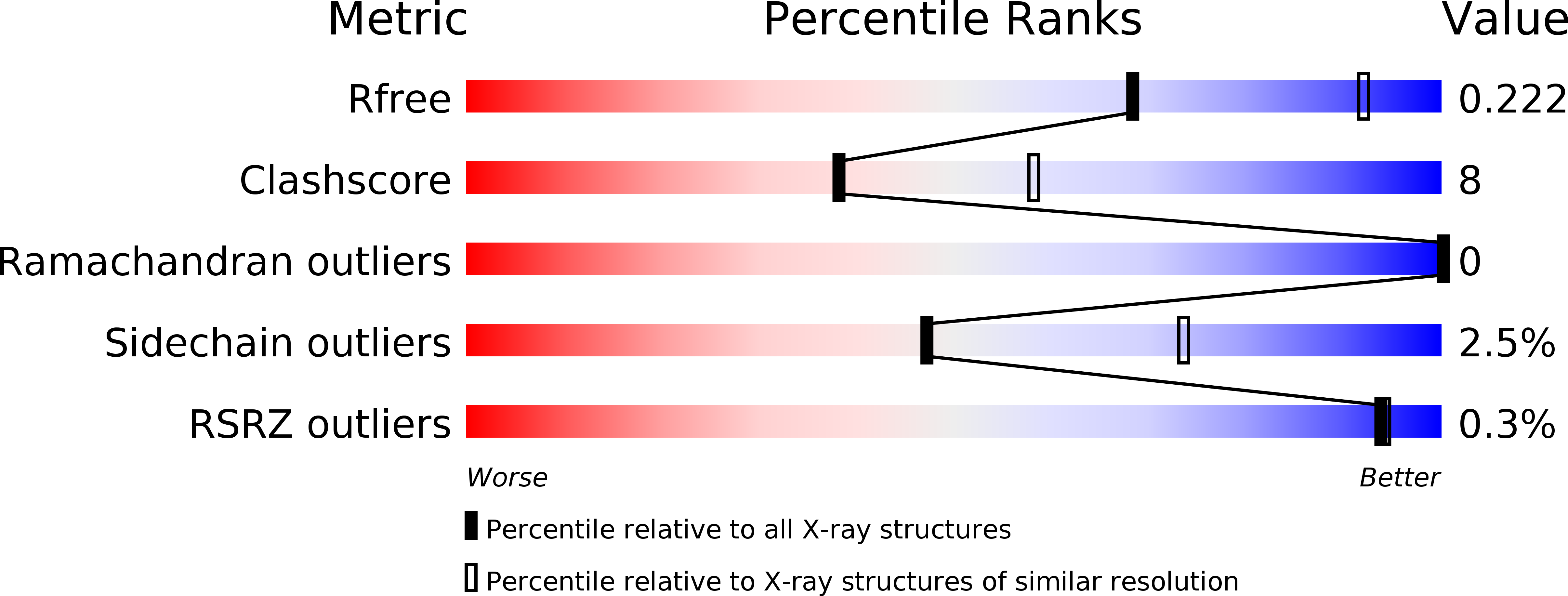
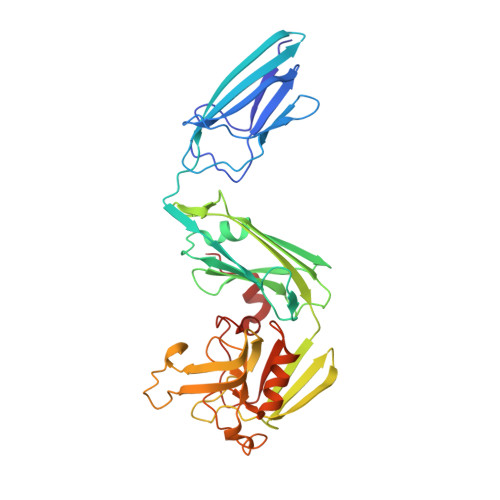
Structural insight into the inactivation of Mycobacterium tuberculosis non-classical transpeptidase LdtMt2 by biapenem and tebipenem.
Bianchet, M.A., Pan, Y.H., Basta, L.A.B., Saavedra, H., Lloyd, E.P., Kumar, P., Mattoo, R., Townsend, C.A., Lamichhane, G.(2017) BMC Biochem 18: 8-8
- PubMed: 28545389
- DOI: https://doi.org/10.1186/s12858-017-0082-4
- Primary Citation of Related Structures:
5D7H, 5DC2, 5DCC - PubMed Abstract:
The carbapenem subclass of β-lactams is among the most potent antibiotics available today. Emerging evidence shows that, unlike other subclasses of β-lactams, carbapenems bind to and inhibit non-classical transpeptidases (L,D-transpeptidases) that generate 3 → 3 linkages in bacterial peptidoglycan. The carbapenems biapenem and tebipenem exhibit therapeutically valuable potencies against Mycobacterium tuberculosis (Mtb). Here, we report the X-ray crystal structures of Mtb L,D-transpeptidase-2 (Ldt Mt2 ) complexed with biapenem or tebipenem. Despite significant variations in carbapenem sulfur side chains, biapenem and tebipenem ultimately form an identical adduct that docks to the outer cavity of Ldt Mt2 . We propose that this common adduct is an enzyme catalyzed decomposition of the carbapenem adduct by a mechanism similar to S-conjugate elimination by β-lyases. The results presented here demonstrate biapenem and tebipenem bind to the outer cavity of Ldt Mt2 , covalently inactivate the enzyme, and subsequently degrade via an S-conjugate elimination mechanism. We discuss structure based drug design based on the findings and propose that the S-conjugate elimination can be leveraged to design novel agents to deliver and locally release antimicrobial factors to act synergistically with the carbapenem carrier.
Organizational Affiliation:
Department of Neurology, Johns Hopkins University School of Medicine, 725 N. Wolfe Street, Baltimore, MD, 21205, USA. [email protected].