Seeding for sirtuins: microseed matrix seeding to obtain crystals of human Sirt3 and Sirt2 suitable for soaking.
Rumpf, T., Gerhardt, S., Einsle, O., Jung, M.(2015) Acta Crystallogr F Struct Biol Commun 71: 1498-1510
- PubMed: 26625292
- DOI: https://doi.org/10.1107/S2053230X15019986
- Primary Citation of Related Structures:
5D7N, 5D7O, 5D7P, 5D7Q - PubMed Abstract:
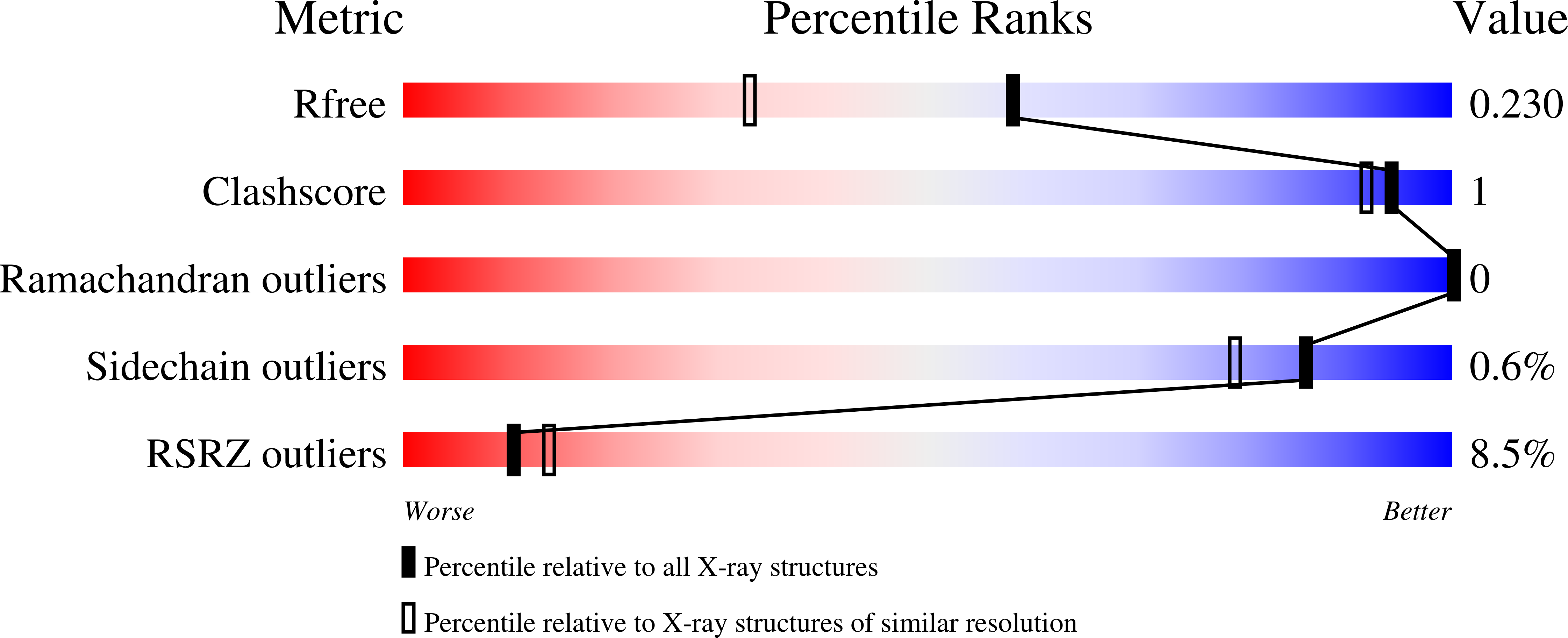
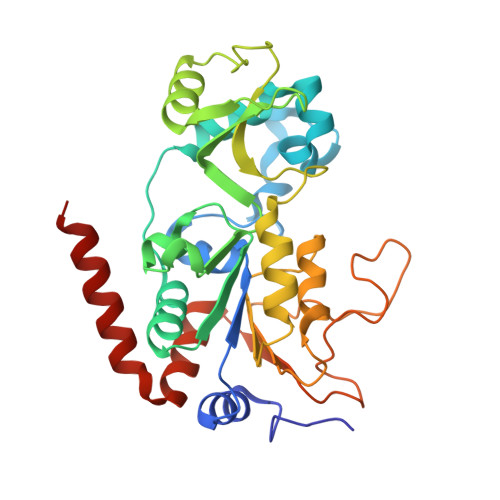
Sirtuins constitute a family of NAD(+)-dependent enzymes that catalyse the cleavage of various acyl groups from the ℇ-amino group of lysines. They regulate a series of cellular processes and their misregulation has been implicated in various diseases, making sirtuins attractive drug targets. To date, only a few sirtuin modulators have been reported that are suitable for cellular research and their development has been hampered by a lack of structural information. In this work, microseed matrix seeding (MMS) was used to obtain crystals of human Sirt3 in its apo form and of human Sirt2 in complex with ADP ribose (ADPR). Crystal formation using MMS was predictable, less error-prone and yielded a higher number of crystals per drop than using conventional crystallization screening methods. The crystals were used to solve the crystal structures of apo Sirt3 and of Sirt2 in complex with ADPR at an improved resolution, as well as the crystal structures of Sirt2 in complex with ADPR and the indoles EX527 and CHIC35. These Sirt2-ADPR-indole complexes unexpectedly contain two indole molecules and provide novel insights into selective Sirt2 inhibition. The MMS approach for Sirt2 and Sirt3 may be used as the basis for structure-based optimization of Sirt2/3 inhibitors in the future.
Organizational Affiliation:
Institute of Pharmaceutical Sciences, Albert-Ludwigs-University Freiburg, Albertstrasse 25, 79104 Freiburg, Baden-Württemberg, Germany.