Discovery of Potent, Selective, and Orally Bioavailable Small-Molecule Modulators of the Mediator Complex-Associated Kinases CDK8 and CDK19.
Mallinger, A., Schiemann, K., Rink, C., Stieber, F., Calderini, M., Crumpler, S., Stubbs, M., Adeniji-Popoola, O., Poeschke, O., Busch, M., Czodrowski, P., Musil, D., Schwarz, D., Ortiz-Ruiz, M.J., Schneider, R., Thai, C., Valenti, M., de Haven Brandon, A., Burke, R., Workman, P., Dale, T., Wienke, D., Clarke, P.A., Esdar, C., Raynaud, F.I., Eccles, S.A., Rohdich, F., Blagg, J.(2016) J Med Chem 59: 1078-1101
- PubMed: 26796641
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01685
- Primary Citation of Related Structures:
5FGK, 5HBE, 5HBH, 5HBJ - PubMed Abstract:
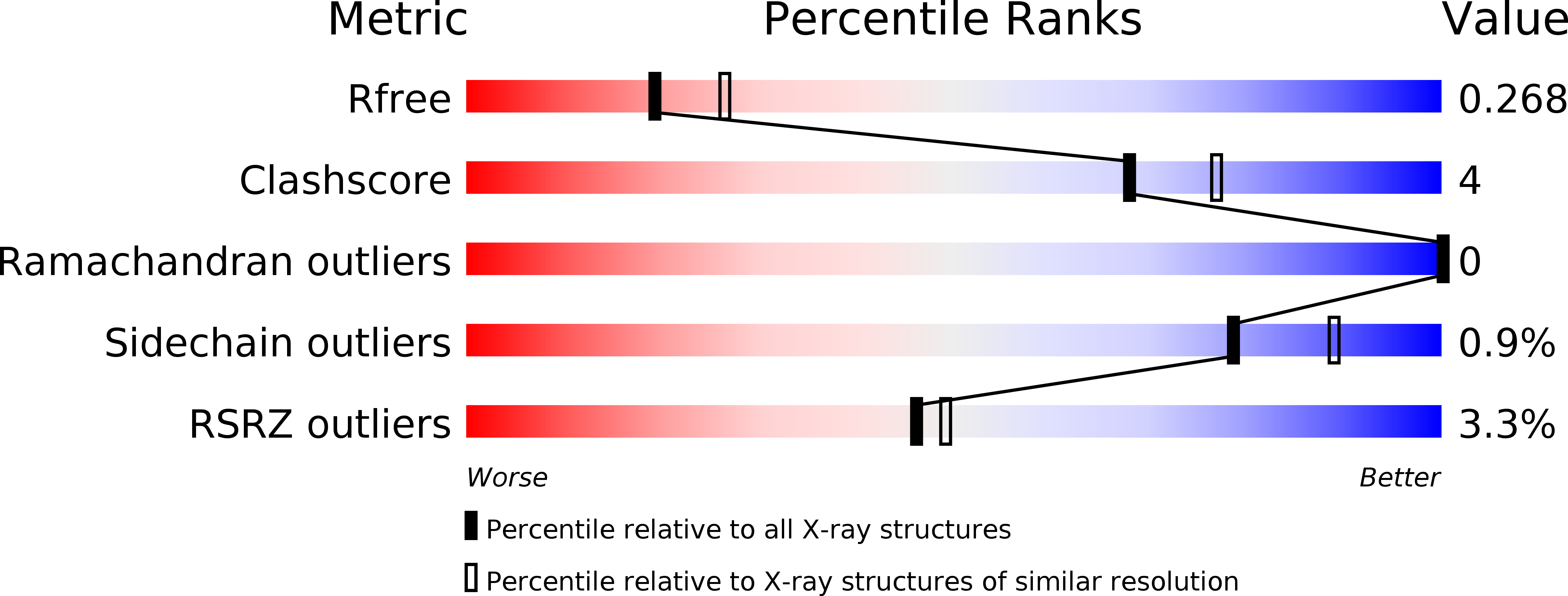
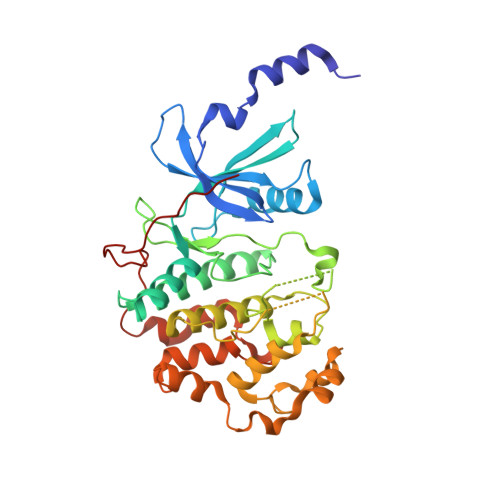
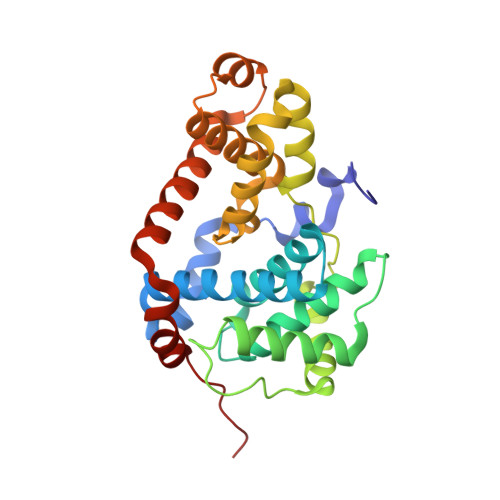
The Mediator complex-associated cyclin-dependent kinase CDK8 has been implicated in human disease, particularly in colorectal cancer where it has been reported as a putative oncogene. Here we report the discovery of 109 (CCT251921), a potent, selective, and orally bioavailable inhibitor of CDK8 with equipotent affinity for CDK19. We describe a structure-based design approach leading to the discovery of a 3,4,5-trisubstituted-2-aminopyridine series and present the application of physicochemical property analyses to successfully reduce in vivo metabolic clearance, minimize transporter-mediated biliary elimination while maintaining acceptable aqueous solubility. Compound 109 affords the optimal compromise of in vitro biochemical, pharmacokinetic, and physicochemical properties and is suitable for progression to animal models of cancer.
Organizational Affiliation:
Cancer Research UK Cancer Therapeutics Unit at The Institute of Cancer Research, London, SW7 3RP, U.K.