New Insights on Signal Propagation by Sensory Rhodopsin II/Transducer Complex.
Ishchenko, A., Round, E., Borshchevskiy, V., Grudinin, S., Gushchin, I., Klare, J.P., Remeeva, A., Polovinkin, V., Utrobin, P., Balandin, T., Engelhard, M., Buldt, G., Gordeliy, V.(2017) Sci Rep 7: 41811-41811
- PubMed: 28165484
- DOI: https://doi.org/10.1038/srep41811
- Primary Citation of Related Structures:
5JJE, 5JJF, 5JJJ, 5JJN - PubMed Abstract:
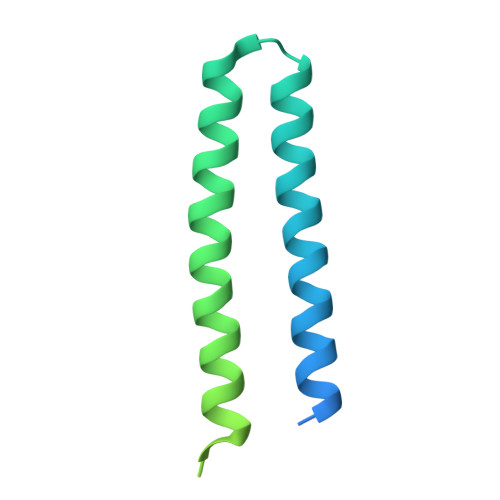
The complex of two membrane proteins, sensory rhodopsin II (NpSRII) with its cognate transducer (NpHtrII), mediates negative phototaxis in halobacteria N. pharaonis. Upon light activation NpSRII triggers a signal transduction chain homologous to the two-component system in eubacterial chemotaxis. Here we report on crystal structures of the ground and active M-state of the complex in the space group I2 1 2 1 2 1 . We demonstrate that the relative orientation of symmetrical parts of the dimer is parallel ("U"-shaped) contrary to the gusset-like ("V"-shaped) form of the previously reported structures of the NpSRII/NpHtrII complex in the space group P2 1 2 1 2, although the structures of the monomers taken individually are nearly the same. Computer modeling of the HAMP domain in the obtained "V"- and "U"-shaped structures revealed that only the "U"-shaped conformation allows for tight interactions of the receptor with the HAMP domain. This is in line with existing data and supports biological relevance of the "U" shape in the ground state. We suggest that the "V"-shaped structure may correspond to the active state of the complex and transition from the "U" to the "V"-shape of the receptor-transducer complex can be involved in signal transduction from the receptor to the signaling domain of NpHtrII.
Organizational Affiliation:
Institute of Complex Systems (ICS), ICS-6: Structural Biochemistry, Research Centre Jülich, 52425 Jülich, Germany.