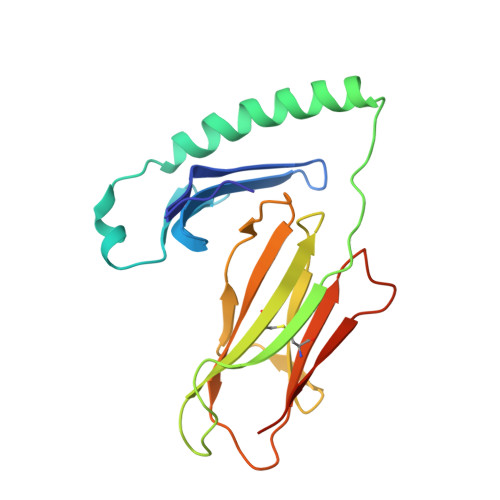
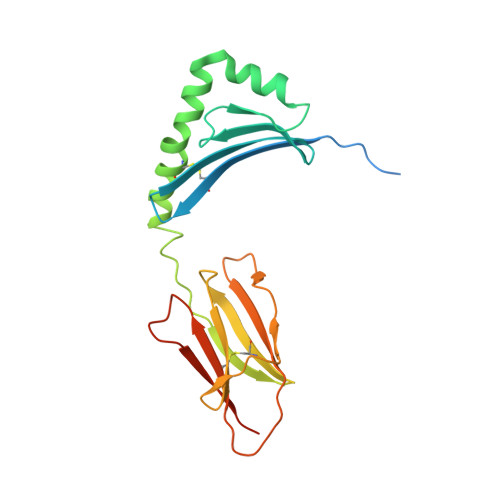
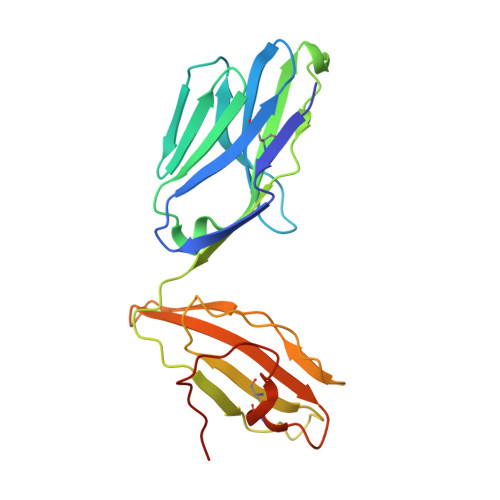
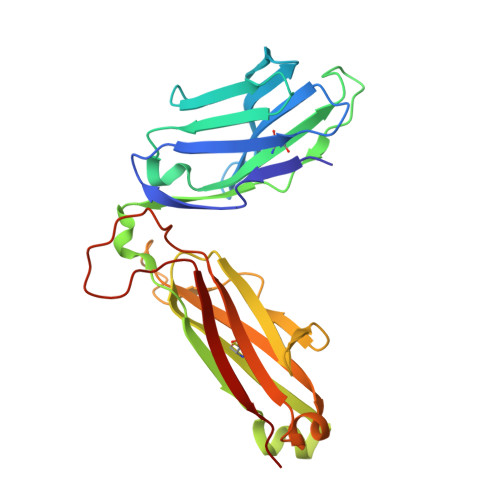
Diverse T Cell Receptor Gene Usage in HLA-DQ8-Associated Celiac Disease Converges into a Consensus Binding Solution.
Petersen, J., Kooy-Winkelaar, Y., Loh, K.L., Tran, M., van Bergen, J., Koning, F., Rossjohn, J., Reid, H.H.(2016) Structure 24: 1643-1657
- PubMed: 27568928
- DOI: https://doi.org/10.1016/j.str.2016.07.010
- Primary Citation of Related Structures:
5KS9, 5KSA, 5KSB - PubMed Abstract:
In HLA-DQ8-associated celiac disease, TRAV26-2 + -TRBV9 + and TRAV8-3 + -TRBV6 + T cells recognize the immunodominant DQ8-glia-α1 epitope, whereupon a non-germline-encoded arginine residue played a key role in binding HLA-DQ8-glia-α1. Whether distinct T cell receptor (TCR) recognition modes exist for gliadin epitopes remains unclear. TCR repertoire analysis revealed populations of HLA-DQ8-glia-α1 and HLA-DQ8.5-glia-γ1 restricted TRAV20 + -TRBV9 + T cells that did not possess a non-germline-encoded arginine residue. The crystal structures of a TRAV20 + -TRBV9 + TCR-HLA-DQ8-glia-α1 complex and two TRAV20 + -TRBV9 + TCR-HLA-DQ8.5-glia-γ1 complexes were determined. This revealed that the differential specificity toward DQ8-glia-α1 and DQ8.5-glia-γ1 was governed by CDR3β-loop-mediated interactions. Surprisingly, a germline-encoded arginine residue within the CDR1α loop of the TRAV20 + TCR substituted for the role of the non-germline-encoded arginine in the TRAV26-2 + -TRBV9 + and TRAV8-3 + -TRBV6 + TCRs. Thus, in celiac disease, the responding TCR repertoire is driven by a common mechanism that selects for structural elements within the TCR that have convergent binding solutions in HLA-DQ8-gliadin recognition.
Organizational Affiliation:
Infection and Immunity Program, The Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, VIC 3800, Australia; Australian Research Council Centre of Excellence in Advanced Molecular Imaging, Monash University, Clayton, VIC 3800, Australia.