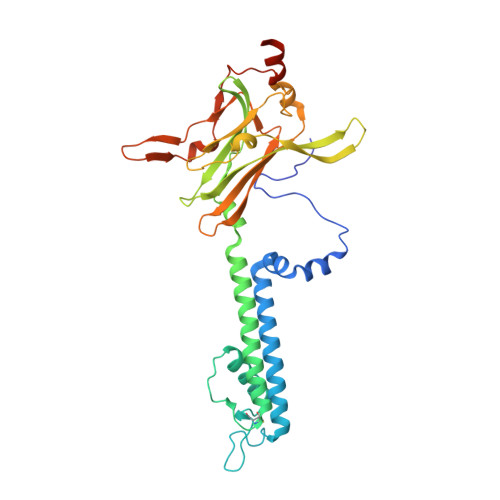
Structural basis of control of inward rectifier Kir2 channel gating by bulk anionic phospholipids.
Lee, S.J., Ren, F., Zangerl-Plessl, E.M., Heyman, S., Stary-Weinzinger, A., Yuan, P., Nichols, C.G.(2016) J Gen Physiol 148: 227-237
- PubMed: 27527100
- DOI: https://doi.org/10.1085/jgp.201611616
- Primary Citation of Related Structures:
5KUK, 5KUM - PubMed Abstract:
Inward rectifier potassium (Kir) channel activity is controlled by plasma membrane lipids. Phosphatidylinositol-4,5-bisphosphate (PIP2) binding to a primary site is required for opening of classic inward rectifier Kir2.1 and Kir2.2 channels, but interaction of bulk anionic phospholipid (PL(-)) with a distinct second site is required for high PIP2 sensitivity. Here we show that introduction of a lipid-partitioning tryptophan at the second site (K62W) generates high PIP2 sensitivity, even in the absence of PL(-) Furthermore, high-resolution x-ray crystal structures of Kir2.2[K62W], with or without added PIP2 (2.8- and 2.0-Å resolution, respectively), reveal tight tethering of the C-terminal domain (CTD) to the transmembrane domain (TMD) in each condition. Our results suggest a refined model for phospholipid gating in which PL(-) binding at the second site pulls the CTD toward the membrane, inducing the formation of the high-affinity primary PIP2 site and explaining the positive allostery between PL(-) binding and PIP2 sensitivity.
Organizational Affiliation:
Department of Cell Biology and Physiology, Washington University School of Medicine, St. Louis, MO 63110 Center for the Investigation of Membrane Excitability Diseases, Washington University School of Medicine, St. Louis, MO 63110.