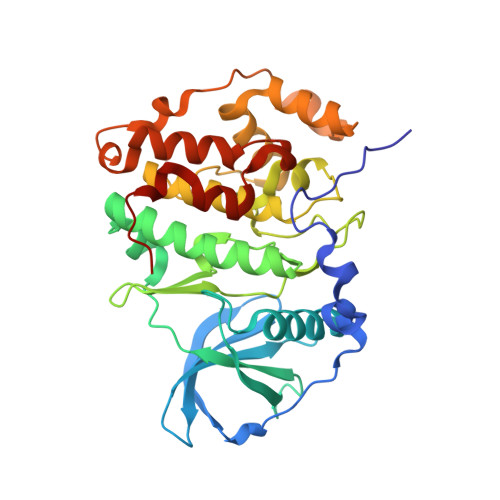
A pi-Halogen Bond of Dibenzofuranones with the Gatekeeper Phe113 in Human Protein Kinase CK2 Leads to Potent Tight Binding Inhibitors.
Schnitzler, A., Gratz, A., Bollacke, A., Weyrich, M., Kucklander, U., Wunsch, B., Gotz, C., Niefind, K., Jose, J.(2018) Pharmaceuticals (Basel) 11
- PubMed: 29462988
- DOI: https://doi.org/10.3390/ph11010023
- Primary Citation of Related Structures:
5N9K, 5N9L, 5N9N - PubMed Abstract:
Human protein kinase CK2 is an emerging target for neoplastic diseases. Potent lead structures for human CK2 inhibitors are derived from dibenzofuranones. Two new derivatives, 7,9-dichloro-1,2-dihydro-8-hydroxy-4-[(4-methoxyphenylamino)-methylene]dibenzo[ b , d ]furan-3(2 H )-one ( 4a ) and ( E )-1,3-dichloro-6-[(4-methoxyphenylimino)-methyl]dibenzo[ b , d ]furan-2,7-diol ( 5 ) were tested for inhibition of CK2 and induction of apoptosis in LNCaP cells. Both turned out to be tight binding inhibitors, with IC 50 values of 7 nM ( 4a ) and 5 nM ( 5 ) and an apparent K i value of 0.4 nM for both. Compounds 4a and 5 reduced cellular CK2 activity, indicating cell permeability. Cell viability was substantially impaired in LNCaP cells, as well as apoptosis was induced, which was not appearing in non-neoplastic ARPE-19 cells. Co-crystallization of 4a and 5 revealed an unexpected π -halogen bond of the chloro substituent at C9 with the gatekeeper amino acid Phe113, leading to an inverted binding mode in comparison to parent compound 4b , with the Cl at C6 instead, which was co-crystallized as a control. This indicates that the position of the chloro substituent on ring A of the dibenzofuran scaffold is responsible for an inversion of the binding mode that enhances potency.
Organizational Affiliation:
Institut für Biochemie, Department für Chemie, Universität zu Köln, Zülpicher Straße 47, D-50674 Köln, Germany. [email protected].