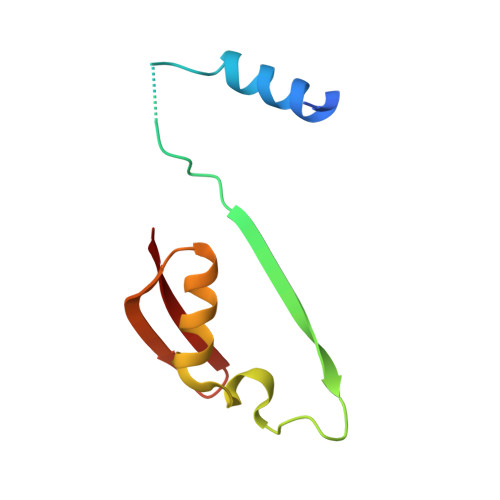
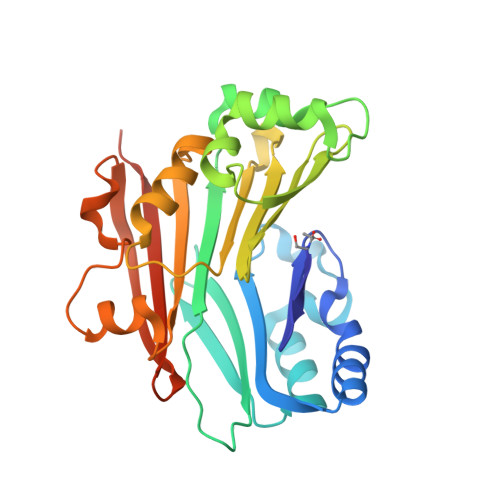
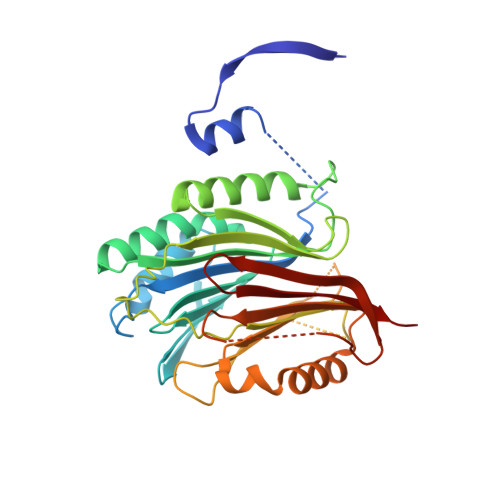
Relief of autoinhibition by conformational switch explains enzyme activation by a catalytically dead paralog.
Volkov, O.A., Kinch, L., Ariagno, C., Deng, X., Zhong, S., Grishin, N., Tomchick, D.R., Chen, Z., Phillips, M.A.(2016) Elife 5
- PubMed: 27977001
- DOI: https://doi.org/10.7554/eLife.20198
- Primary Citation of Related Structures:
5TVF, 5TVM, 5TVO - PubMed Abstract:
Catalytically inactive enzyme paralogs occur in many genomes. Some regulate their active counterparts but the structural principles of this regulation remain largely unknown. We report X-ray structures of Trypanosoma brucei S -adenosylmethionine decarboxylase alone and in functional complex with its catalytically dead paralogous partner, prozyme. We show monomeric Tb AdoMetDC is inactive because of autoinhibition by its N-terminal sequence. Heterodimerization with prozyme displaces this sequence from the active site through a complex mechanism involving a cis -to- trans proline isomerization, reorganization of a β-sheet, and insertion of the N-terminal α-helix into the heterodimer interface, leading to enzyme activation. We propose that the evolution of this intricate regulatory mechanism was facilitated by the acquisition of the dimerization domain, a single step that can in principle account for the divergence of regulatory schemes in the AdoMetDC enzyme family. These studies elucidate an allosteric mechanism in an enzyme and a plausible scheme by which such complex cooperativity evolved.
Organizational Affiliation:
Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, United States.