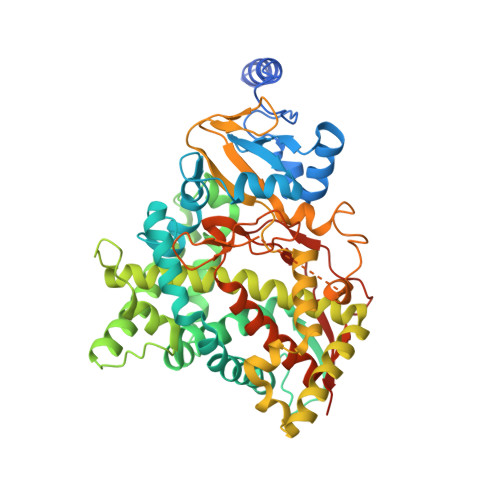
Crystal Structures of Full-Length Lanosterol 14 alpha-Demethylases of Prominent Fungal Pathogens Candida albicans and Candida glabrata Provide Tools for Antifungal Discovery.
Keniya, M.V., Sabherwal, M., Wilson, R.K., Woods, M.A., Sagatova, A.A., Tyndall, J.D.A., Monk, B.C.(2018) Antimicrob Agents Chemother 62
- PubMed: 30126961
- DOI: https://doi.org/10.1128/AAC.01134-18
- Primary Citation of Related Structures:
5JLC, 5V5Z - PubMed Abstract:
Targeting lanosterol 14α-demethylase (LDM) with azole drugs provides prophylaxis and treatments for superficial and disseminated fungal infections, but cure rates are not optimal for immunocompromised patients and individuals with comorbidities. The efficacy of azole drugs has also been reduced due to the emergence of drug-resistant fungal pathogens. We have addressed the need to improve the potency, spectrum, and specificity for azoles by expressing in Saccharomyces cerevisiae functional, recombinant, hexahistidine-tagged, full-length Candida albicans LDM (CaLDM6×His) and Candida glabrata LDM (CgLDM6×His) and determining their X-ray crystal structures. The crystal structures of CaLDM6×His, CgLDM6×His, and ScLDM6×His have the same fold and bind itraconazole in nearly identical conformations. The catalytic domains of the full-length LDMs have the same fold as the CaLDM6×His catalytic domain in complex with posaconazole, with minor structural differences within the ligand binding pocket. Our structures give insight into the LDM reaction mechanism and phenotypes of single-site CaLDM mutations. This study provides a practical basis for the structure-directed discovery of novel antifungals that target LDMs of fungal pathogens.
Organizational Affiliation:
Sir John Walsh Research Institute, Faculty of Dentistry, University of Otago, Dunedin, New Zealand.