Structural and mutational analyses of psychrophilic and mesophilic adenylate kinases highlight the role of hydrophobic interactions in protein thermal stability.
Moon, S., Kim, J., Koo, J., Bae, E.(2019) Struct Dyn 6: 024702-024702
- PubMed: 31111079
- DOI: https://doi.org/10.1063/1.5089707
- Primary Citation of Related Structures:
5X6I, 5X6J - PubMed Abstract:
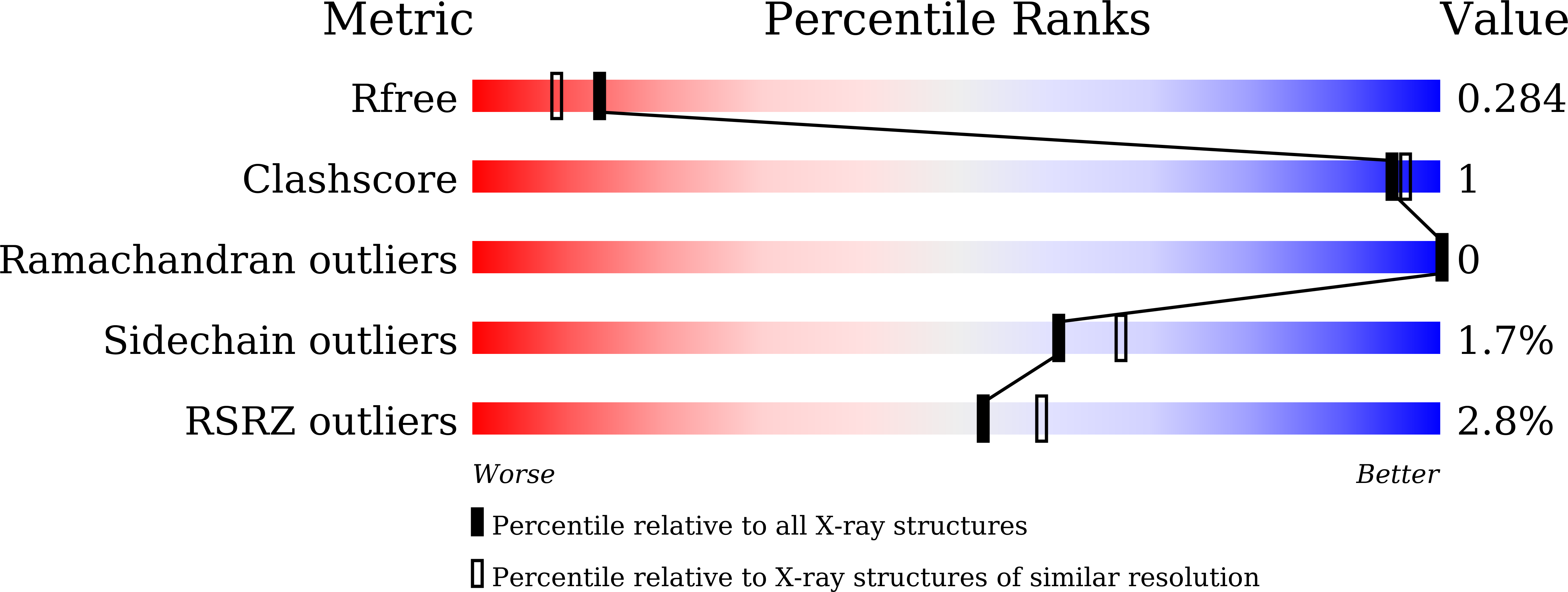
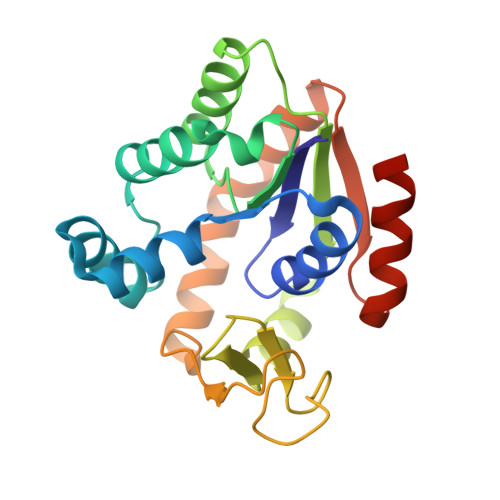
Protein thermal stability is an important field since thermally stable proteins are desirable in many academic and industrial settings. Information on protein thermal stabilization can be obtained by comparing homologous proteins from organisms living at distinct temperatures. Here, we report structural and mutational analyses of adenylate kinases (AKs) from psychrophilic Bacillus globisporus (AKp) and mesophilic Bacillus subtilis (AKm). Sequence and structural comparison showed suboptimal hydrophobic packing around Thr26 in the CORE domain of AKp, which was replaced with an Ile residue in AKm. Mutations that improved hydrophobicity of the Thr residue increased the thermal stability of the psychrophilic AKp, and the largest stabilization was observed for a Thr-to-Ile substitution. Furthermore, a reverse Ile-to-Thr mutation in the mesophilic AKm significantly decreased thermal stability. We determined the crystal structures of mutant AKs to confirm the impact of the residue substitutions on the overall stability. Taken together, our results provide a structural basis for the stability difference between psychrophilic and mesophilic AK homologues and highlight the role of hydrophobic interactions in protein thermal stability.
Organizational Affiliation:
Department of Agricultural Biotechnology, Seoul National University, Seoul 08826, South Korea.