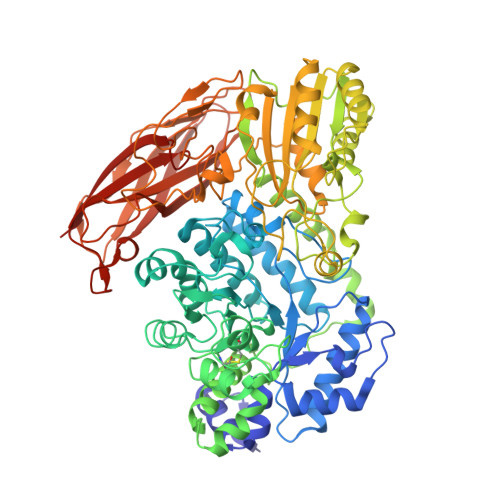
Crystal structure and substrate recognition mechanism of Aspergillus oryzae isoprimeverose-producing enzyme.
Matsuzawa, T., Watanabe, M., Nakamichi, Y., Fujimoto, Z., Yaoi, K.(2019) J Struct Biol 205: 84-90
- PubMed: 30445155
- DOI: https://doi.org/10.1016/j.jsb.2018.11.005
- Primary Citation of Related Structures:
5YOT, 5YQS - PubMed Abstract:
Isoprimeverose-producing enzymes (IPases) release isoprimeverose (α-d-xylopyranosyl-(1 → 6)-d-glucopyranose) from the non-reducing end of xyloglucan oligosaccharides. Aspergillus oryzae IPase (IpeA) is classified as a member of the glycoside hydrolase family 3 (GH3); however, it has unusual substrate specificity compared with other GH3 enzymes. Xylopyranosyl branching at the non-reducing ends of xyloglucan oligosaccharides is vital for IpeA activity. We solved the crystal structure of IpeA with isoprimeverose at 2.4 Å resolution, showing that the structure of IpeA formed a dimer and was composed of three domains: an N-terminal (β/α) 8 TIM-barrel domain, α/β/α sandwich fold domain, and a C-terminal fibronectin-like domain. The catalytic TIM-barrel domain possessed a catalytic nucleophile (Asp300) and acid/base (Glu524) residues. Interestingly, we found that the cavity of the active site of IpeA was larger than that of other GH3 enzymes, and subsite -1' played an important role in its activity. The glucopyranosyl and xylopyranosyl residues of isoprimeverose were located at subsites -1 and -1', respectively. Gln58 and Tyr89 contributed to the interaction with the xylopyranosyl residue of isoprimeverose through hydrogen bonding and stacking effects, respectively. Our findings provide new insights into the substrate recognition of GH3 enzymes.
Organizational Affiliation:
Bioproduction Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Tsukuba Central 6, 1-1-1 Higashi, Tsukuba, Ibaraki 305-8566, Japan.