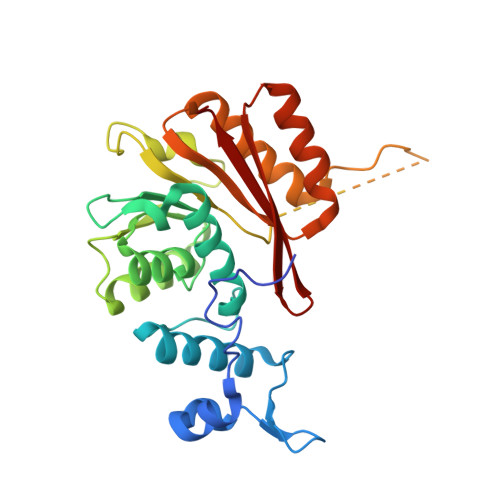
Structural insights into the RNA methyltransferase domain of METTL16.
Ruszkowska, A., Ruszkowski, M., Dauter, Z., Brown, J.A.(2018) Sci Rep 8: 5311-5311
- PubMed: 29593291
- DOI: https://doi.org/10.1038/s41598-018-23608-8
- Primary Citation of Related Structures:
6B91, 6B92 - PubMed Abstract:
N 6 -methyladenosine (m 6 A) is an abundant modification in messenger RNA and noncoding RNAs that affects RNA metabolism. Methyltransferase-like protein 16 (METTL16) is a recently confirmed m 6 A RNA methyltransferase that methylates U6 spliceosomal RNA and interacts with the 3'-terminal RNA triple helix of MALAT1 (metastasis-associated lung adenocarcinoma transcript 1). Here, we present two X-ray crystal structures of the N-terminal methyltransferase domain (residues 1-291) of human METTL16 (METTL16_291): an apo structure at 1.9 Å resolution and a post-catalytic S-adenosylhomocysteine-bound complex at 2.1 Å resolution. The structures revealed a highly conserved Rossmann fold that is characteristic of Class I S-adenosylmethionine-dependent methyltransferases and a large, positively charged groove. This groove likely represents the RNA-binding site and it includes structural elements unique to METTL16. In-depth analysis of the active site led to a model of the methyl transfer reaction catalyzed by METTL16. In contrast to the major m 6 A methyltransferase heterodimer METTL3/METTL14, full-length METTL16 forms a homodimer and METTL16_291 exists as a monomer based on size-exclusion chromatography. A native gel-shift assay shows that METTL16 binds to the MALAT1 RNA triple helix, but monomeric METTL16_291 does not. Our results provide insights into the molecular structure of METTL16, which is distinct from METTL3/METTL14.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, IN, 46556, USA.