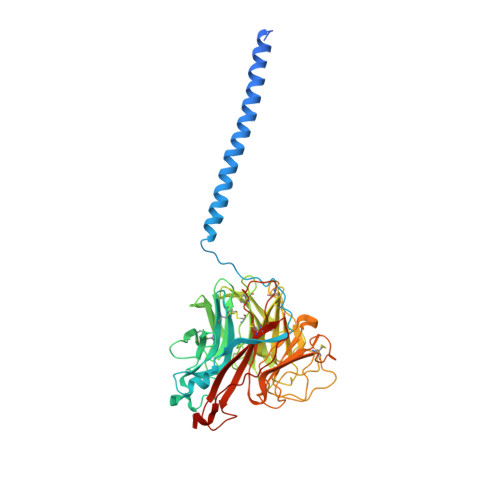
Structure of an Influenza A virus N9 neuraminidase with a tetrabrachion-domain stalk.
Streltsov, V.A., Schmidt, P.M., McKimm-Breschkin, J.L.(2019) Acta Crystallogr F Struct Biol Commun 75: 89-97
- PubMed: 30713159
- DOI: https://doi.org/10.1107/S2053230X18017892
- Primary Citation of Related Structures:
6CRD, 6D3B, 6MCX - PubMed Abstract:
The influenza neuraminidase (NA) is a homotetramer with head, stalk, transmembrane and cytoplasmic regions. The structure of the NA head with a stalk has never been determined. The NA head from an N9 subtype influenza A virus, A/tern/Australia/G70C/1975 (H1N9), was expressed with an artificial stalk derived from the tetrabrachion (TB) tetramerization domain from Staphylothermus marinus. The NA was successfully crystallized both with and without the TB stalk, and the structures were determined to 2.6 and 2.3 Å resolution, respectively. Comparisons of the two NAs with the native N9 NA structure from egg-grown virus showed that the artificial TB stalk maintained the native NA head structure, supporting previous biological observations.
Organizational Affiliation:
CSIRO Manufacturing, 343 Royal Parade, Parkville, Victoria 3052, Australia.