Azole Resistance Reduces Susceptibility to the Tetrazole Antifungal VT-1161.
Monk, B.C., Keniya, M.V., Sabherwal, M., Wilson, R.K., Graham, D.O., Hassan, H.F., Chen, D., Tyndall, J.D.A.(2019) Antimicrob Agents Chemother 63
- PubMed: 30397057
- DOI: https://doi.org/10.1128/AAC.02114-18
- Primary Citation of Related Structures:
5UL0, 6E8Q - PubMed Abstract:
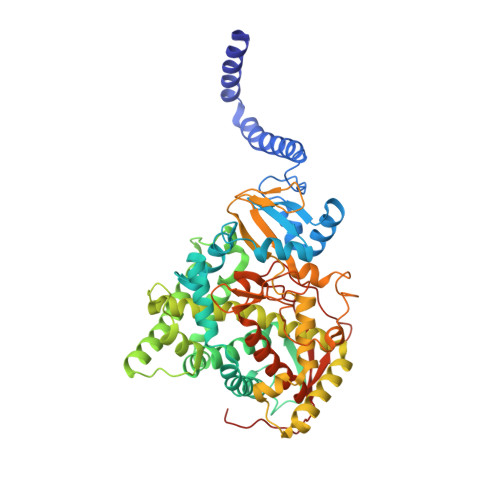
Tetrazole antifungals designed to target fungal lanosterol 14α-demethylase (LDM) appear to be effective against a range of fungal pathogens. In addition, a crystal structure of the catalytic domain of Candida albicans LDM in complex with the tetrazole VT-1161 has been obtained. We have addressed concern about artifacts that might arise from crystallizing VT-1161 with truncated recombinant CYP51s and measured the impact on VT-1161 susceptibility of genotypes known to confer azole resistance. A yeast system was used to overexpress recombinant full-length Saccharomyces cerevisiae LDM with a C-terminal hexahistidine tag (ScLDM6×His) for phenotypic analysis and crystallographic studies with VT-1161 or with the widely used triazole drug posaconazole (PCZ). We determined the effect of characterized mutations in LDM on VT-1161 activity and identified drug efflux pumps from fungi, including key fungal pathogens, that efflux VT-1161. The relevance of these yeast-based observations on drug efflux was verified using clinical isolates of C. albicans and Candida glabrata VT-1161 binding elicits a significant conformational difference between the full-length and truncated enzymes not found when posaconazole is bound. Susceptibility to VT-1161 is reduced by ATP-binding cassette (ABC) and major facilitator superfamily (MFS) drug efflux pumps, the overexpression of LDM, and mutations within the drug binding pocket of LDM that affect interaction with the tertiary alcohol of the drug.
Organizational Affiliation:
Sir John Walsh Research Institute, Faculty of Dentistry, University of Otago, Dunedin, New Zealand [email protected].